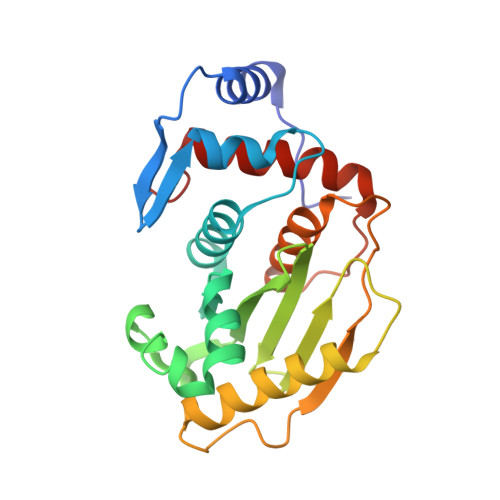
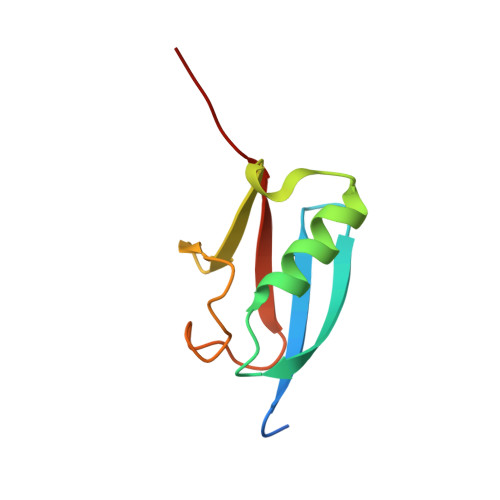
Ulp1-SUMO crystal structure and genetic analysis reveal conserved interactions and a regulatory element essential for cell growth in yeast.
Mossessova, E., Lima, C.D.(2000) Mol Cell 5: 865-876
- PubMed: 10882122 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(00)80326-3
- Primary Citation Related Structures:
1EUV - PubMed Abstract:
Modification of cellular proteins by the ubiquitin-like protein SUMO is essential for nuclear processes and cell cycle progression in yeast. The Ulp1 protease catalyzes two essential functions in the SUMO pathway: (1) processing of full-length SUMO to its mature form and (2) deconjugation of SUMO from targeted proteins. Selective reduction of the proteolytic reaction produced a covalent thiohemiacetal transition state complex between a Ulp1 C-terminal fragment and its cellular substrate Smt3, the yeast SUMO homolog. The Ulp1-Smt3 crystal structure and functional testing of elements within the conserved interface elucidate determinants of SUMO recognition, processing, and deconjugation. Genetic analysis guided by the structure further reveals a regulatory element N-terminal to the proteolytic domain that is required for cell growth in yeast.
- Biochemistry Department, Weill Medical College of Cornell University, New York, New York 10021, USA.
Organizational Affiliation: