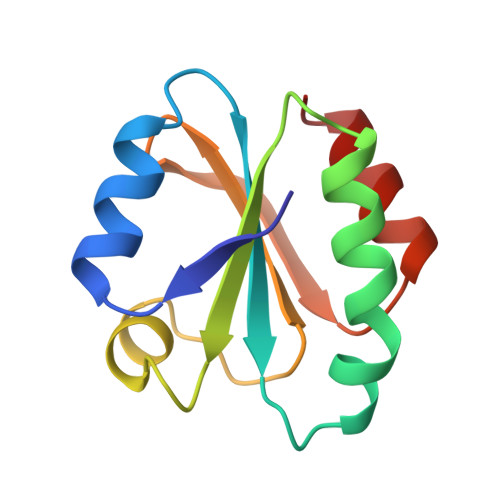
Crystal structures of reduced, oxidized, and mutated human thioredoxins: evidence for a regulatory homodimer.
Weichsel, A., Gasdaska, J.R., Powis, G., Montfort, W.R.(1996) Structure 4: 735-751
- PubMed: 8805557 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(96)00079-2
- Primary Citation Related Structures:
1ERT, 1ERU, 1ERV, 1ERW - PubMed Abstract:
Human thioredoxin reduces the disulfide bonds of numerous proteins in vitro, and can activate transcription factors such as NFkB in vivo. Thioredoxin can also act as a growth factor, and is overexpressed and secreted in certain tumor cells. Crystal structures were determined for reduced and oxidized wild type human thioredoxin (at 1.7 and 2.1 A nominal resolution, respectively), and for reduced mutant proteins Cys73-->Ser and Cys32-->Ser/Cys35-->Ser (at 1.65 and 1.8 A, respectively). Surprisingly, thioredoxin is dimeric in all four structures; the dimer is linked through a disulfide bond between Cys73 of each monomer, except in Cys73-->Ser where a hydrogen bond occurs. The thioredoxin active site is blocked by dimer formation. Conformational changes in the active site and dimer interface accompany oxidation of the active-site cysteines, Cys32 and Cys35. It has been suggested that a reduced pKa in the first cysteine (Cys32 in human thioredoxin) of the active-site sequence is important for modulation of the redox potential in thioredoxin. A hydrogen bond between the sulfhydryls of Cys32 and Cys35 may reduce the pKa of Cys32 and this pKa depression probably results in increased nucleophilicity of the Cys32 thiolate group. This nucleophilicity, in tum, is thought to be necessary for the role of thioredoxin in disulfide-bond reduction. The physiological role, if any, of thioredoxin dimer formation remains unknown. It is possible that dimerization may provide a mechanism for regulation of the protein, or a means of sensing oxidative stress.
- Department of Biochemistry, University of Arizona, Tucson 85721, USA.
Organizational Affiliation: