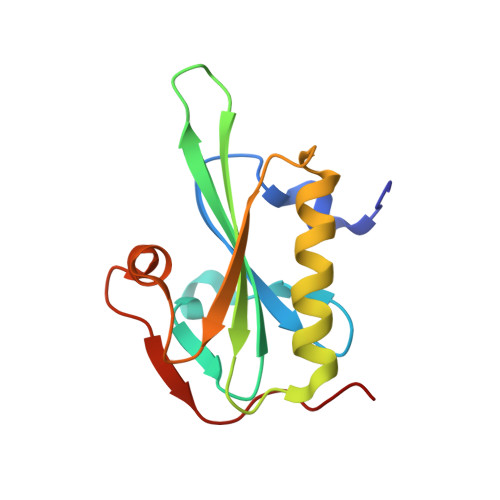
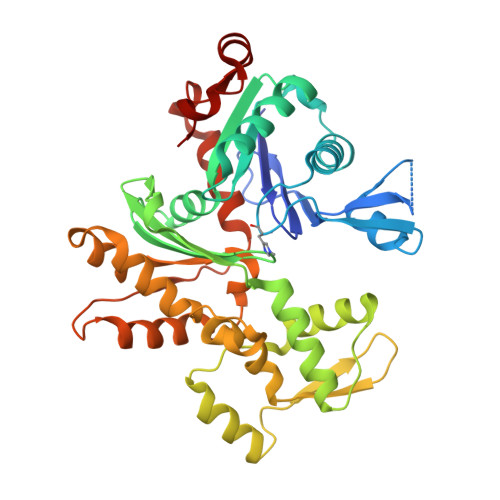
Structure of gelsolin segment 1-actin complex and the mechanism of filament severing.
McLaughlin, P.J., Gooch, J.T., Mannherz, H.G., Weeds, A.G.(1993) Nature 364: 685-692
- PubMed: 8395021 Search on PubMed
- DOI: https://doi.org/10.1038/364685a0
- Primary Citation Related Structures:
1EQY - PubMed Abstract:
The structure of the segment 1 domain of gelsolin, a protein that fragments actin filaments in cells, is reported in complex with actin. Segment 1 binds monomer using an apolar patch rimmed by hydrogen bonds in a cleft between actin domains. On the actin filament model it binds tangentially, disrupting only those contacts between adjacent subunits in one helical strand. The segment 1 fold is general for all segments of the gelsolin family because the conserved residues form the core of the structure. It also provides a basis for understanding the origin of an amyloidosis caused by a gelsolin variant.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: