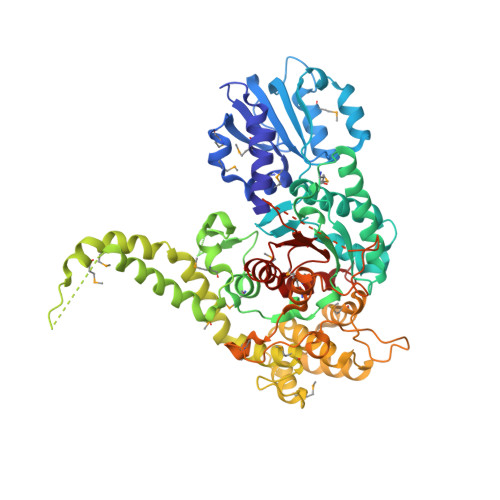
The X-ray crystal structure of neuronal Sec1 from squid sheds new light on the role of this protein in exocytosis.
Bracher, A., Perrakis, A., Dresbach, T., Betz, H., Weissenhorn, W.(2000) Structure 8: 685-694
- PubMed: 10903948 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00156-8
- Primary Citation Related Structures:
1EPU - PubMed Abstract:
Sec1-like molecules have been implicated in a variety of eukaryotic vesicle transport processes including neurotransmitter release by exocytosis. They regulate vesicle transport by binding to a t-SNARE from the syntaxin family. This process is thought to prevent SNARE complex formation, a protein complex required for membrane fusion. Whereas Sec1 molecules are essential for neurotransmitter release and other secretory events, their interaction with syntaxin molecules seems to represent a negative regulatory step in secretion. Here we report the X-ray crystal structure of a neuronal Sec1 homologue from squid, s-Sec1, at 2.4 A resolution. Neuronal s-Sec1 is a modular protein that folds into a V-shaped three-domain assembly. Peptide and mutagenesis studies are discussed with respect to the mechanism of Sec1 regulation. Comparison of the structure of squid s-Sec1 with the previously determined structure of rat neuronal Sec1 (n-Sec1) bound to syntaxin-1a indicates conformational rearrangements in domain III induced by syntaxin binding. The crystal structure of s-Sec1 provides the molecular scaffold for a number of molecular interactions that have been reported to affect Sec1 function. The structural differences observed between s-Sec1 and the structure of a rat n-Sec1-syntaxin-1a complex suggest that local conformational changes are sufficient to release syntaxin-1a from neuronal Sec1, an active process that is thought to involve additional effector molecule(s).
- European Molecular Biology Laboratory (EMBL), Grenoble, 38000, France.
Organizational Affiliation: