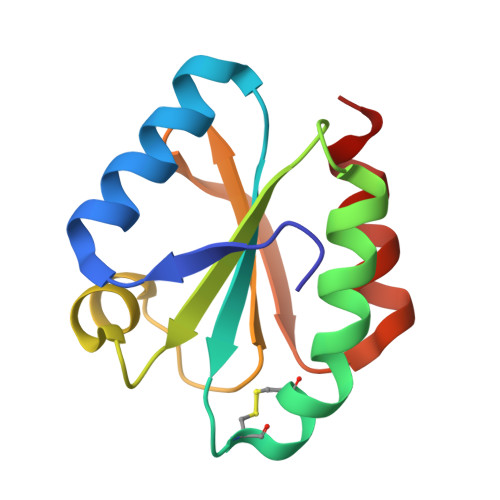
Crystal structure of the wild-type and D30A mutant thioredoxin h of Chlamydomonas reinhardtii and implications for the catalytic mechanism.
Menchise, V., Corbier, C., Didierjean, C., Saviano, M., Benedetti, E., Jacquot, J.P., Aubry, A.(2001) Biochem J 359: 65-75
- PubMed: 11563970 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/0264-6021:3590065
- Primary Citation Related Structures:
1EP7, 1EP8 - PubMed Abstract:
Thioredoxins are ubiquitous proteins which catalyse the reduction of disulphide bridges on target proteins. The catalytic mechanism proceeds via a mixed disulphide intermediate whose breakdown should be enhanced by the involvement of a conserved buried residue, Asp-30, as a base catalyst towards residue Cys-39. We report here the crystal structure of wild-type and D30A mutant thioredoxin h from Chlamydomonas reinhardtii, which constitutes the first crystal structure of a cytosolic thioredoxin isolated from a eukaryotic plant organism. The role of residue Asp-30 in catalysis has been revisited since the distance between the carboxylate OD1 of Asp-30 and the sulphur SG of Cys-39 is too great to support the hypothesis of direct proton transfer. A careful analysis of all available crystal structures reveals that the relative positioning of residues Asp-30 and Cys-39 as well as hydrophobic contacts in the vicinity of residue Asp-30 do not allow a conformational change sufficient to bring the two residues close enough for a direct proton transfer. This suggests that protonation/deprotonation of Cys-39 should be mediated by a water molecule. Molecular-dynamics simulations, carried out either in vacuo or in water, as well as proton-inventory experiments, support this hypothesis. The results are discussed with respect to biochemical and structural data.
- Laboratoire de Cristallographie et Modélisation des Matériaux Minéraux et Biologiques, Groupe Biocristallographie, ESA 7036, Université Henri Poincaré-Nancy I, BP 239, 54506 Vandoeuvre-lès-Nancy Cedex, France.
Organizational Affiliation: