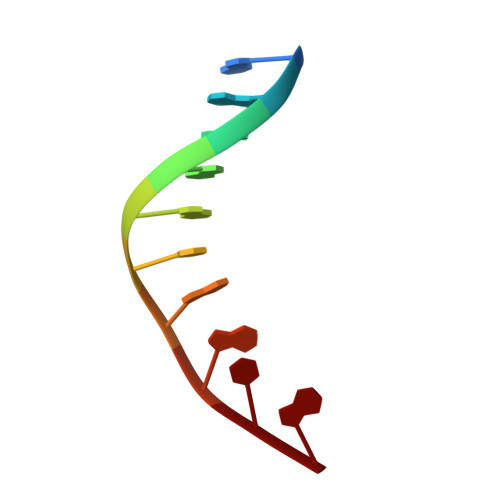
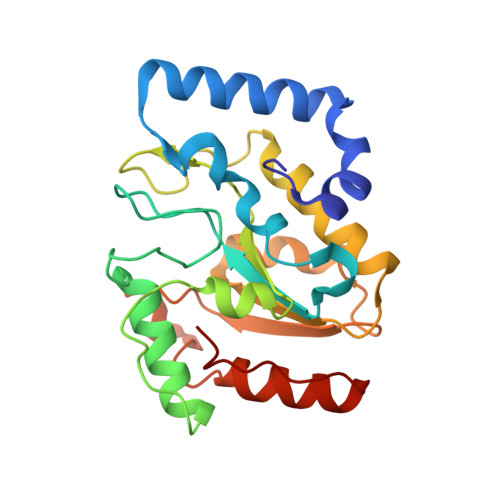
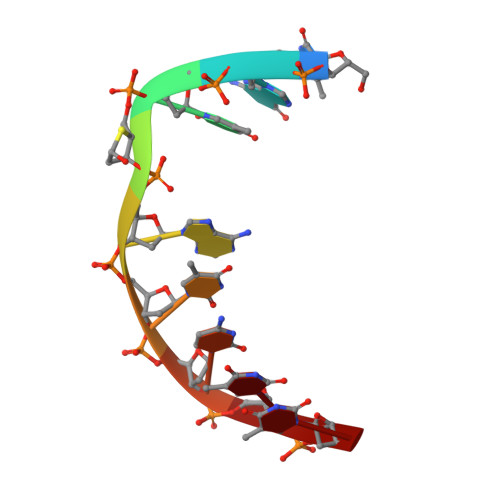
Uracil-DNA glycosylase-DNA substrate and product structures: conformational strain promotes catalytic efficiency by coupled stereoelectronic effects.
Parikh, S.S., Walcher, G., Jones, G.D., Slupphaug, G., Krokan, H.E., Blackburn, G.M., Tainer, J.A.(2000) Proc Natl Acad Sci U S A 97: 5083-5088
- PubMed: 10805771 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.97.10.5083
- Primary Citation Related Structures:
1EMH, 1EMJ - PubMed Abstract:
Enzymatic transformations of macromolecular substrates such as DNA repair enzyme/DNA transformations are commonly interpreted primarily by active-site functional-group chemistry that ignores their extensive interfaces. Yet human uracil-DNA glycosylase (UDG), an archetypical enzyme that initiates DNA base-excision repair, efficiently excises the damaged base uracil resulting from cytosine deamination even when active-site functional groups are deleted by mutagenesis. The 1.8-A resolution substrate analogue and 2.0-A resolution cleaved product cocrystal structures of UDG bound to double-stranded DNA suggest enzyme-DNA substrate-binding energy from the macromolecular interface is funneled into catalytic power at the active site. The architecturally stabilized closing of UDG enforces distortions of the uracil and deoxyribose in the flipped-out nucleotide substrate that are relieved by glycosylic bond cleavage in the product complex. This experimentally defined substrate stereochemistry implies the enzyme alters the orientation of three orthogonal electron orbitals to favor electron transpositions for glycosylic bond cleavage. By revealing the coupling of this anomeric effect to a delocalization of the glycosylic bond electrons into the uracil aromatic system, this structurally implicated mechanism resolves apparent paradoxes concerning the transpositions of electrons among orthogonal orbitals and the retention of catalytic efficiency despite mutational removal of active-site functional groups. These UDG/DNA structures and their implied dissociative excision chemistry suggest biology favors a chemistry for base-excision repair initiation that optimizes pathway coordination by product binding to avoid the release of cytotoxic and mutagenic intermediates. Similar excision chemistry may apply to other biological reaction pathways requiring the coordination of complex multistep chemical transformations.
- Skaggs Institute for Chemical Biology and the Department of Molecular Biology, MB-4, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037-1027, USA.
Organizational Affiliation: