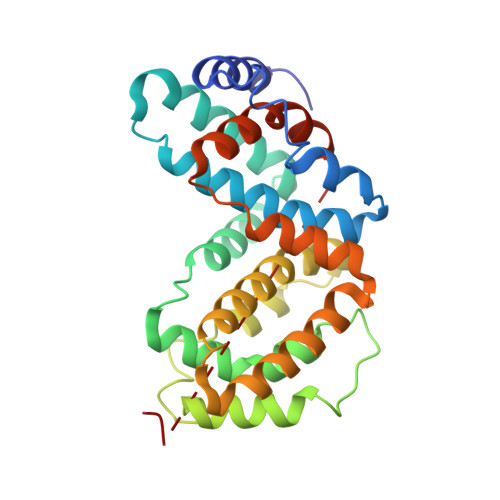
Design, characterization, and structure of a biologically active single-chain mutant of human IFN-gamma.
Landar, A., Curry, B., Parker, M.H., DiGiacomo, R., Indelicato, S.R., Nagabhushan, T.L., Rizzi, G., Walter, M.R.(2000) J Mol Biology 299: 169-179
- PubMed: 10860730 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3734
- Primary Citation Related Structures:
1EKU - PubMed Abstract:
A mutant form of human interferon-gamma (IFN-gamma SC1) that binds one IFN-gamma receptor alpha chain (IFN-gamma R alpha) has been designed and characterized. IFN-gamma SC1 was derived by linking the two peptide chains of the IFN-gamma dimer by a seven-residue linker and changing His111 in the first chain to an aspartic acid residue. Isothermal titration calorimetry shows that IFN-gamma SC1 forms a 1:1 complex with its high-affinity receptor (IFN-gamma R alpha) with an affinity of 27(+/- 9) nM. The crystal structure of IFN-gamma SC1 has been determined at 2.9 A resolution from crystals grown in 1.4 M citrate solutions at pH 7.6. Comparison of the wild-type receptor-binding domain and the Asp111-containing domain of IFN-gamma SC1 show that they are structurally equivalent but have very different electrostatic surface potentials. As a result, surface charge rather than structural changes is likely responsible for the inability of the His111-->Asp domain of to bind IFN-gamma R alpha. The AB loops of IFN-gamma SC1 adopt conformations similar to the ordered loops of IFN-gamma observed in the crystal structure of the IFN-gamma/IFN-gamma R alpha complex. Thus, IFN-gamma R alpha binding does not result in a large conformational change in the AB loop as previously suggested. The structure also reveals the final six C-terminal amino acid residues of IFN-gamma SC1 (residues 253-258) that have not been observed in any other reported IFN-gamma structures. Despite binding to only one IFN-gamma R alpha, IFN-gamma SC1 is biologically active in cell proliferation, MHC class I induction, and anti-viral assays. This suggests that one domain of IFN-gamma is sufficient to recruit IFN-gamma R alpha and IFN-gamma R beta into a complex competent for eliciting biological activity. The current data are consistent with the main role of the IFN-gamma dimer being to decrease the dissociation constant of IFN-gamma for its cellular receptors.
- Center for Macromolecular Crystallography, University of Alabama, Birmingham 35294, USA.
Organizational Affiliation: