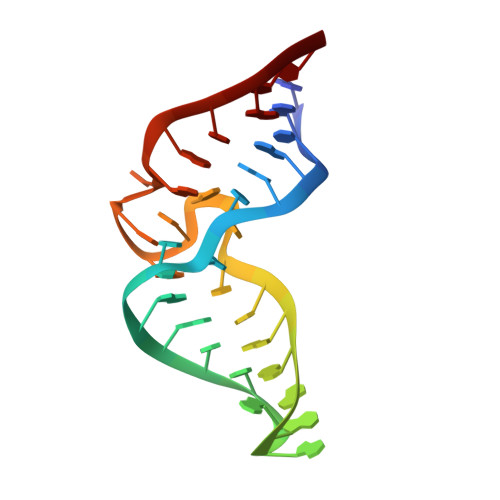
Interlocking structural motifs mediate molecular discrimination by a theophylline-binding RNA.
Zimmermann, G.R., Jenison, R.D., Wick, C.L., Simorre, J.P., Pardi, A.(1997) Nat Struct Biol 4: 644-649
- PubMed: 9253414 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0897-644
- Primary Citation Related Structures:
1EHT - PubMed Abstract:
To visualize the interplay of RNA structural interactions in a ligand binding site, we have determined the solution structure of a high affinity RNA-theophylline complex using NMR spectroscopy. The structure provides insight into the ability of this in vitro selected RNA to discriminate theophylline from the structurally similar molecule caffeine. Numerous RNA structural motifs combine to form a well-ordered binding pocket where an intricate network of hydrogen bonds and stacking interactions lock the theophylline into the complex. Two internal loops interact to form the binding site which consists of a sandwich of three base triples. The complex also contains novel base-zipper and 1-3-2 stacking motifs, in addition to an adenosine platform and a reversed sugar. An important feature of the RNA is that many of the conserved core residues participate in multiple overlapping tertiary interactions. This complex illustrates how interlocking structural motifs can be assembled into a highly specific ligand-binding site that possesses high levels of affinity and molecular discrimination.
- Department of Chemistry and Biochemistry, University of Colorado, Boulder 80309-0215, USA.
Organizational Affiliation: