Sequence-specific 1H n.m.r. assignments and determination of the three-dimensional structure of reduced Escherichia coli glutaredoxin.
Sodano, P., Xia, T.H., Bushweller, J.H., Bjornberg, O., Holmgren, A., Billeter, M., Wuthrich, K.(1991) J Mol Biology 221: 1311-1324
- PubMed: 1942053
- DOI: https://doi.org/10.1016/0022-2836(91)90935-y
- Primary Citation of Related Structures:
1EGR - PubMed Abstract:
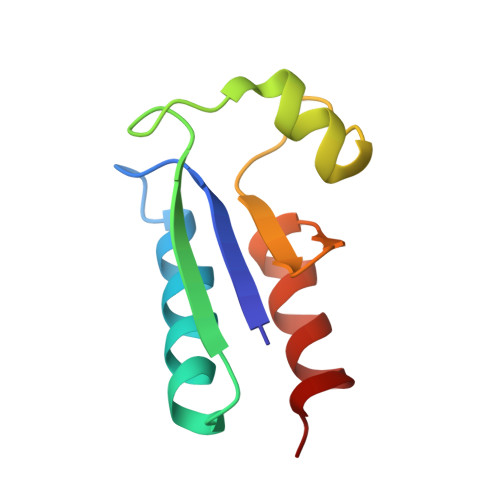
The determination of the nuclear magnetic resonance structure of reduced E. coli glutaredoxin in aqueous solution is described. Based on nearly complete, sequence-specific resonance assignments, 813 nuclear Overhauser effect distance constraints and 191 dihedral angle constraints were employed as the input for the structure calculations, for which the distance geometry program DIANA was used followed by simulated annealing with the program X-PLOR. The molecular architecture of reduced glutaredoxin is made up of three helices and four-stranded beta-sheet. The first strand of the beta-sheet (residues 2 to 7) runs parallel to the second strand (32 to 37) and antiparallel to the third strand (61 to 64), and the sheet is extended in an antiparallel fashion with a fourth strand (67 to 69). The first helix with residues 13 to 28 and the last helix (71 to 83) run parallel to each other on one side of the beta-sheet, with their direction opposite to that of the two parallel beta-strands, and the helix formed by residues 44 to 53 fills space available due to the twist of the beta-sheet and the reduced length of the last two beta-strands. The active site Cys11-Pro-Tyr-Cys14 is located after the first beta-strand and occupies the latter part of the loop connecting this strand with the first helix.
- Institut für Molekularbiologie und Biophysik, Eidgenössiche Technische Hochschule-Hönggerberg, Zürich, Switzerland.
Organizational Affiliation: