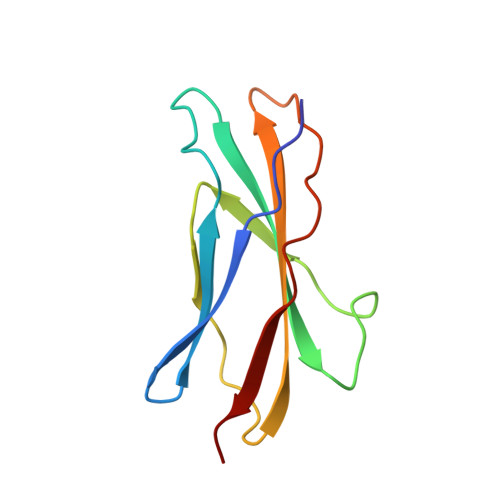
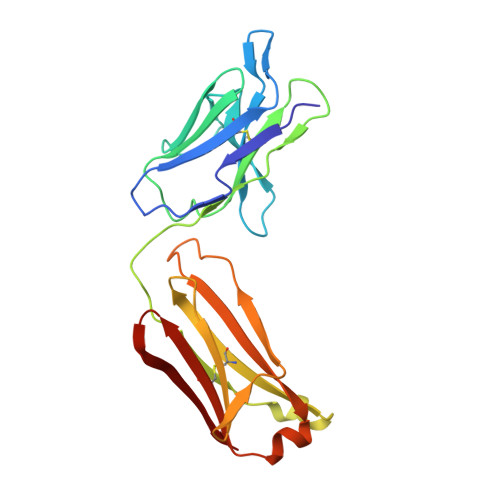
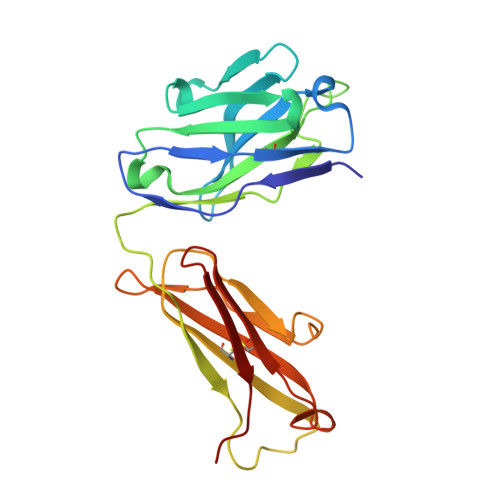
Structure of the activation domain of the GM-CSF/IL-3/IL-5 receptor common beta-chain bound to an antagonist.
Rossjohn, J., McKinstry, W.J., Woodcock, J.M., McClure, B.J., Hercus, T.R., Parker, M.W., Lopez, A.F., Bagley, C.J.(2000) Blood 95: 2491-2498
- PubMed: 10753826 Search on PubMed
- Primary Citation Related Structures:
1EGJ - PubMed Abstract:
Heterodimeric cytokine receptors generally consist of a major cytokine-binding subunit and a signaling subunit. The latter can transduce signals by more than 1 cytokine, as exemplified by the granulocyte-macrophage colony-stimulating factor (GM-CSF), interleukin-2 (IL-2), and IL-6 receptor systems. However, often the signaling subunits in isolation are unable to bind cytokines, a fact that has made it more difficult to obtain structural definition of their ligand-binding sites. This report details the crystal structure of the ligand-binding domain of the GM-CSF/IL-3/IL-5 receptor beta-chain (beta(c)) signaling subunit in complex with the Fab fragment of the antagonistic monoclonal antibody, BION-1. This is the first single antagonist of all 3 known eosinophil-producing cytokines, and it is therefore capable of regulating eosinophil-related diseases such as asthma. The structure reveals a fibronectin type III domain, and the antagonist-binding site involves major contributions from the loop between the B and C strands and overlaps the cytokine-binding site. Furthermore, tyrosine(421) (Tyr(421)), a key residue involved in receptor activation, lies in the neighboring loop between the F and G strands, although it is not immediately adjacent to the cytokine-binding residues in the B-C loop. Interestingly, functional experiments using receptors mutated across these loops demonstrate that they are cooperatively involved in full receptor activation. The experiments, however, reveal subtle differences between the B-C loop and Tyr(421), which is suggestive of distinct functional roles. The elucidation of the structure of the ligand-binding domain of beta(c) also suggests how different cytokines recognize a single receptor subunit, which may have implications for homologous receptor systems. (Blood. 2000;95:2491-2498)
- Ian Potter Foundation Protein Crystallography Laboratory, St. Vincent's Institute of Medical Research, Fitzroy, Victoria, Australia.
Organizational Affiliation: