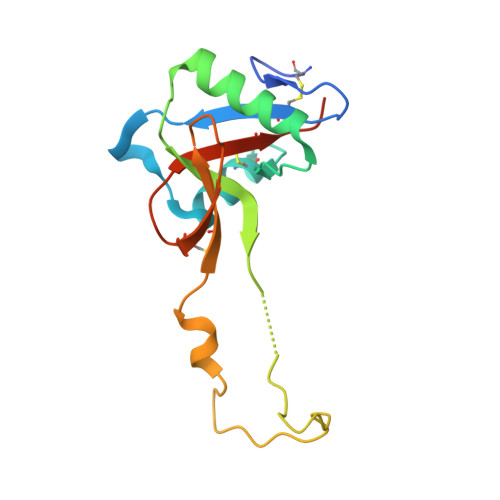
Structure of a C-type carbohydrate recognition domain from the macrophage mannose receptor.
Feinberg, H., Park-Snyder, S., Kolatkar, A.R., Heise, C.T., Taylor, M.E., Weis, W.I.(2000) J Biological Chem 275: 21539-21548
- PubMed: 10779515 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M002366200
- Primary Citation Related Structures:
1EGG, 1EGI - PubMed Abstract:
The mannose receptor of macrophages and liver endothelium mediates clearance of pathogenic organisms and potentially harmful glycoconjugates. The extracellular portion of the receptor includes eight C-type carbohydrate recognition domains (CRDs), of which one, CRD-4, shows detectable binding to monosaccharide ligands. We have determined the crystal structure of CRD-4. Although the basic C-type lectin fold is preserved, a loop extends away from the core of the domain to form a domain-swapped dimer in the crystal. Of the two Ca(2+) sites, only the principal site known to mediate carbohydrate binding in other C-type lectins is occupied. This site is altered in a way that makes sugar binding impossible in the mode observed in other C-type lectins. The structure is likely to represent an endosomal form of the domain formed when Ca(2+) is lost from the auxiliary calcium site. The structure suggests a mechanism for endosomal ligand release in which the auxiliary calcium site serves as a pH sensor. Acid pH-induced removal of this Ca(2+) results in conformational rearrangements of the receptor, rendering it unable to bind carbohydrate ligands.
- Department of Structural Biology, Stanford University School of Medicine, Stanford, California 94305, USA.
Organizational Affiliation: