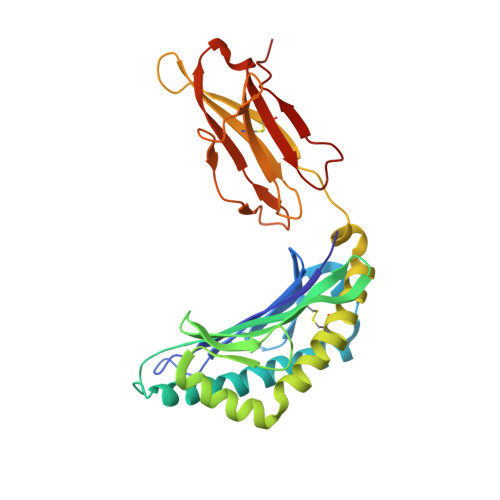
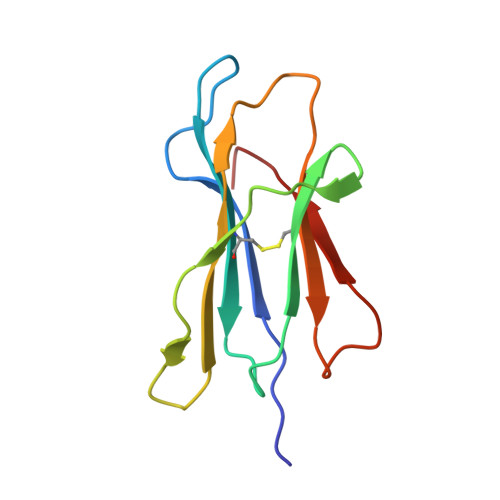
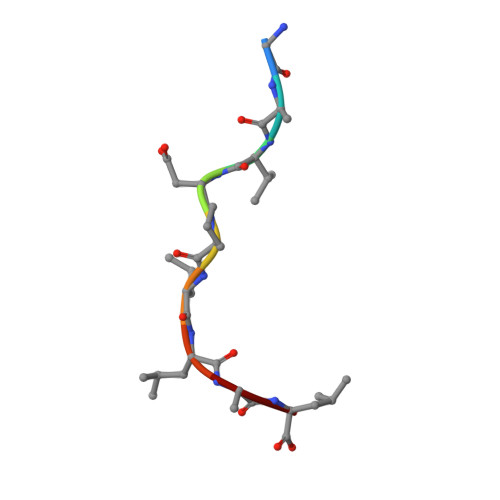
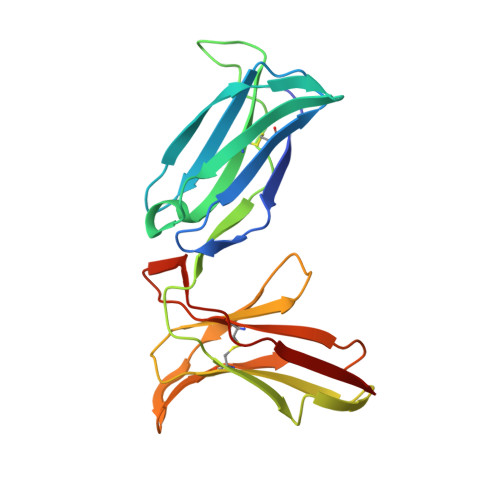
Crystal structure of an NK cell immunoglobulin-like receptor in complex with its class I MHC ligand.
Boyington, J.C., Motyka, S.A., Schuck, P., Brooks, A.G., Sun, P.D.(2000) Nature 405: 537-543
- PubMed: 10850706 Search on PubMed
- DOI: https://doi.org/10.1038/35014520
- Primary Citation Related Structures:
1EFX - PubMed Abstract:
Target cell lysis is regulated by natural killer (NK) cell receptors that recognize class I MHC molecules. Here we report the crystal structure of the human immunoglobulin-like NK cell receptor KIR2DL2 in complex with its class I ligand HLA-Cw3 and peptide. KIR binds in a nearly orthogonal orientation across the alpha1 and alpha2 helices of Cw3 and directly contacts positions 7 and 8 of the peptide. No significant conformational changes in KIR occur on complex formation. The receptor footprint on HLA overlaps with but is distinct from that of the T-cell receptor. Charge complementarity dominates the KIR/HLA interface and mutations that disrupt interface salt bridges substantially diminish binding. Most contacts in the complex are between KIR and conserved HLA-C residues, but a hydrogen bond between Lys 44 of KIR2DL2 and Asn 80 of Cw3 confers the allotype specificity. KIR contact requires position 8 of the peptide to be a residue smaller than valine. A second KIR/HLA interface produced an ordered receptor-ligand aggregation in the crystal which may resemble receptor clustering during immune synapse formation.
- Structural Biology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Rockville, Maryland 20852, USA.
Organizational Affiliation: