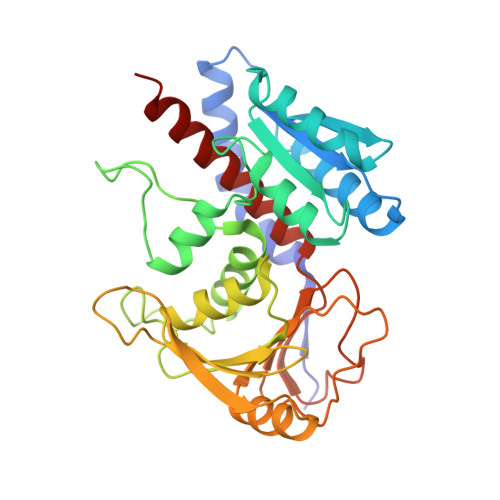
The X-ray structure of the NAD-dependent 5,10-methylenetetrahydrofolate dehydrogenase from Saccharomyces cerevisiae.
Monzingo, A.F., Breksa, A., Ernst, S., Appling, D.R., Robertus, J.D.(2000) Protein Sci 9: 1374-1381
- PubMed: 10933503 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.9.7.1374
- Primary Citation Related Structures:
1EDZ, 1EE9 - PubMed Abstract:
Eucaryotes possess one or more NADP-dependent methylene-THF dehydrogenases as part of multifunctional enzymes. In addition, yeast expresses an unusual monofunctional NAD-dependent enzyme, yMTD. We report X-ray structures for the apoenzyme and its complex with NAD+ at 2.8 and 3.0 A resolution, respectively. The protein fold resembles that seen for the human and Escherichia coli dehydrogenase/cyclohydrolase bifunctional enzymes. The enzyme has two prominent domains, with the active site cleft between them. yMTD has a noncanonical NAD-binding domain that has two inserted strands compared with the NADP-binding domains of the bifunctional enzymes. This insert precludes yMTD from dimerizing in the same way as the bifunctional enzymes. yMTD functions as a dimer, but the mode of dimerization is novel. It does not appear that the difference in dimerization accounts for the difference in cofactor specificity or for the loss of cyclohydrolase activity. These functional differences are probably accounted for by minor differences within the tertiary structure of the active site of the monomeric protein.
- Department of Chemistry and Biochemistry, University of Texas at Austin, 78712, USA.
Organizational Affiliation: