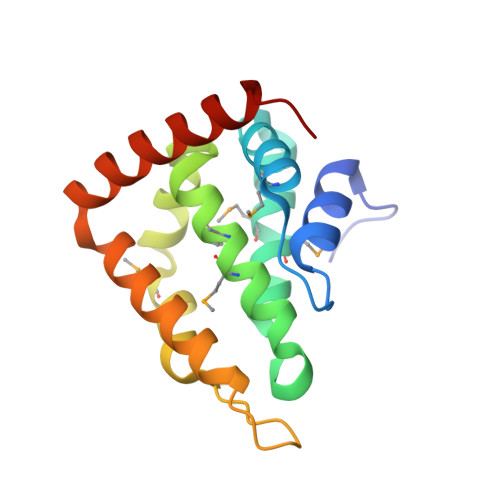
Epsin 1 undergoes nucleocytosolic shuttling and its eps15 interactor NH(2)-terminal homology (ENTH) domain, structurally similar to Armadillo and HEAT repeats, interacts with the transcription factor promyelocytic leukemia Zn(2)+ finger protein (PLZF).
Hyman, J., Chen, H., Di Fiore, P.P., De Camilli, P., Brunger, A.T.(2000) J Cell Biol 149: 537-546
- PubMed: 10791968
- DOI: https://doi.org/10.1083/jcb.149.3.537
- Primary Citation Related Structures:
1EDU - PubMed Abstract:
Epsin (Eps15 interactor) is a cytosolic protein involved in clathrin-mediated endocytosis via its direct interactions with clathrin, the clathrin adaptor AP-2, and Eps15. The NH(2)-terminal portion of epsin contains a phylogenetically conserved module of unknown function, known as the ENTH domain (epsin NH(2)-terminal homology domain). We have now solved the crystal structure of rat epsin 1 ENTH domain to 1.8 A resolution. This domain is structurally similar to armadillo and Heat repeats of beta-catenin and karyopherin-beta, respectively. We have also identified and characterized the interaction of epsin 1, via the ENTH domain, with the transcription factor promyelocytic leukemia Zn(2)+ finger protein (PLZF). Leptomycin B, an antifungal antibiotic, which inhibits the Crm1- dependent nuclear export pathway, induces an accumulation of epsin 1 in the nucleus. These findings suggest that epsin 1 may function in a signaling pathway connecting the endocytic machinery to the regulation of nuclear function.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06520, USA.
Organizational Affiliation: