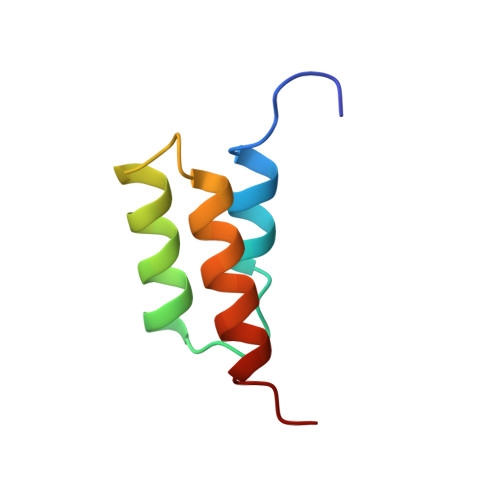
Solution structure of the E-domain of staphylococcal protein A.
Starovasnik, M.A., Skelton, N.J., O'Connell, M.P., Kelley, R.F., Reilly, D., Fairbrother, W.J.(1996) Biochemistry 35: 15558-15569
- PubMed: 8952510
- DOI: https://doi.org/10.1021/bi961409x
- Primary Citation of Related Structures:
1EDI, 1EDJ, 1EDK, 1EDL - PubMed Abstract:
The E-domain of staphylococcal protein A is one of five homologous IgG-binding domains designated E, D, A, B, and C that comprise the extracellular portion of protein A. The E-domain binds tightly to Fc fragments of IgG and binds certain Fv fragments with micromolar affinity. To explore further the structural features of Fc binding by protein A, and as a first step in developing a structural understanding of E-domain/Fv complex formation, we have determined the solution structure of the uncomplexed E-domain using 2D homonuclear and heteronuclear NMR spectroscopy. Complete 1H and 15N resonance assignments were obtained, and the structure was determined from 383 NOE-derived distance restrains, 34 phi and 19 chi 1 dihedral angle restraints, and 54 restraints for 27 H-bonds. 3JH alpha-H beta coupling constants and long-range NOEs involving Phe11 indicate the side chain exists in more than one conformation with differing chi 1 values. NOE restraints that were incompatible with chi 1 = -60 degrees were removed from one set of structure calculations, and those incompatible with chi 1 = 180 degrees were removed from a second set to allow Phe11 to explore both rotamer wells. Thus, two sets of 20 final structures, having no distance or dihedral angle restraint violations greater than 0.12 A or 1.6 degrees, respectively, represent the solution structure of the E-domain. Backbone atomic rms differences with respect to the mean coordinates for each set of 20 structures for residues 8-53 averaged 0.41 +/- 0.06 and 0.35 +/- 0.06 A. No significant differences in the overall structure result from the different orientations of Phe11. The solution structure of the E-domain consists of three alpha-helices that pack together to form a compact helical bundle. A detailed comparison between the E-domain ensembles and the previously determined structure for the B-domain in complex with Fc indicates that only the 180 degrees chi 1 rotamer of Phe11 is competent for binding; the -60 degrees chi 1 rotamer must reorient to 180 degrees to create a cavity that is filled by Ile253 from the CH2 domain of Fc in the Fc-bound complex.
- Department of Protein Engineering, Genentech, Inc., South San Francisco, California 94080, USA.
Organizational Affiliation: