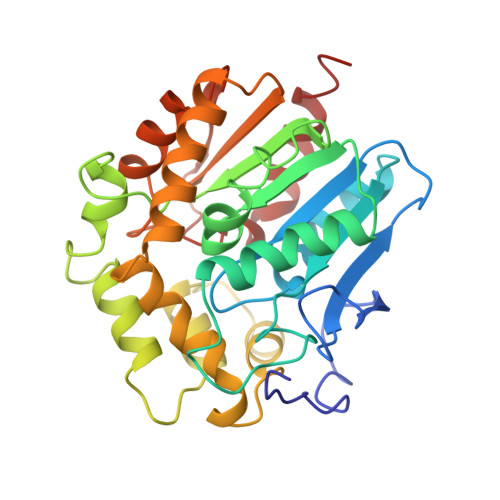
Refined X-ray structures of haloalkane dehalogenase at pH 6.2 and pH 8.2 and implications for the reaction mechanism.
Verschueren, K.H., Franken, S.M., Rozeboom, H.J., Kalk, K.H., Dijkstra, B.W.(1993) J Mol Biology 232: 856-872
- PubMed: 8355275 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1436
- Primary Citation Related Structures:
1EDE - PubMed Abstract:
The crystal structure of haloalkane dehalogenase from Xanthobacter autotrophicus GJ10 has been refined at 1.9 A resolution at two different pH values, the pH of crystallization (pH 6.2) and the pH of optimal activity (pH 8.2), to final R-factors of 16.8% and 16.4%, respectively. Both models show good stereochemical quality. Two non-glycine residues have main-chain torsion angles that are located outside the "allowed" regions in a Ramachandran plot. One of them is the nucleophilic residue Asp124, which, together with the two other active site residues His289 and Asp260, is situated in an internal, predominantly hydrophobic cavity. The other residue, Asn148, helps stabilize the conformations of two of these active-site residues, Asp124 and Asp260. Comparison of the models at pH 6.2 and pH 8.2 revealed one major structural difference. At pH 6.2, a salt-bridge is present between the N epsilon 2 atom of His289 and the O delta 1 atom of Asp124, while at pH 8.2, this salt-bridge is absent, indicating that the N epsilon 2 atom of the histidine residue is mostly deprotonated at the pH of optimum activity. This is in agreement with the putative reaction mechanism in which the O delta 1 atom of Asp124 performs a nucleophilic attack on the substrate, resulting in an intermediate ester. This ester is subsequently cleaved by a hydrolytic water molecule. The high-resolution data sets clearly show the exact position of this water molecule. It is in an ideal position for donating a proton to the N epsilon 2 atom of His289 and subsequently cleaving the covalently bound intermediate ester, releasing the alcohol product. Detailed investigation of both refined models showed a number of unusual structural features. Four out of 11 helices contain an internal proline residue other than in the first turn. Two other alpha-helices have adopted in their central part a 3(10) conformation. A novel four-residue turn between a helix and a strand, the alpha beta 4 turn, is located at the site of the bend in the central eight-stranded beta-sheet of the dehalogenase structure.
- BIOSON Research Institute, University of Groningen, The Netherlands.
Organizational Affiliation: