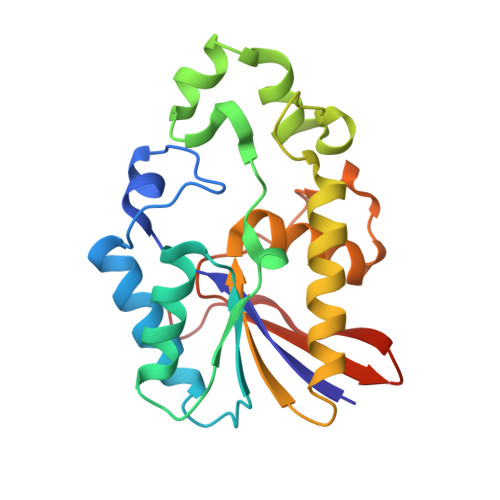
Structure and Mechanism of Action of a Cofactor-Dependent Phosphoglycerate Mutase Homolog from Bacillus Stearothermophilus with Broad Specificity Phosphatase Activity.
Rigden, D.J., Mello, L.V., Setlow, P., Jedrzejas, M.J.(2002) J Mol Biology 315: 1129
- PubMed: 11827481 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5290
- Primary Citation Related Structures:
1EBB - PubMed Abstract:
The crystal structure of Bacillus stearothermophilus PhoE (originally termed YhfR), a broad specificity monomeric phosphatase with a molecular mass of approximately 24 kDa, has been solved at 2.3 A resolution in order to investigate its structure and function. PhoE, already identified as a homolog of a cofactor-dependent phosphoglycerate mutase, shares with the latter an alpha/beta/alpha sandwich structure spanning, as a structural excursion, a smaller subdomain composed of two alpha-helices and one short beta-strand. The active site contains residues from both the alpha/beta/alpha sandwich and the sub-domain. With the exception of the hydrophilic catalytic machinery conserved throughout the cofactor-dependent phosphoglycerate mutase family, the active-site cleft is strikingly hydrophobic. Docking studies with two diverse, favored substrates show that 3-phosphoglycerate may bind to the catalytic core, while alpha-napthylphosphate binding also involves the hydrophobic portion of the active-site cleft. Combining a highly favorable phospho group binding site common to these substrate binding modes and data from related enzymes, a catalytic mechanism can be proposed that involves formation of a phosphohistidine intermediate on His10 and likely acid-base behavior of Glu83. Other structural factors contributing to the broad substrate specificity of PhoE can be identified. The dynamic independence of the subdomain may enable the active-site cleft to accommodate substrates of different sizes, although similar motions are present in simulations of cofactor-dependent phosphoglycerate mutases, perhaps favoring a more general functional role. A significant number of entries in protein sequence databases, particularly from unfinished microbial genomes, are more similar to PhoE than to cofactor-dependent phosphoglycerate mutases or to fructose-2,6-bisphosphatases. This PhoE structure will therefore serve as a valuable basis for inference of structural and functional characteristics of these proteins.
- National Centre of Genetic Resources and Biotechnology, Cenargen/Embrapa, S.A.I.N. Parque Rural, Final W5, Asa Norte, Brasília, 70770-900, Brazil.
Organizational Affiliation: