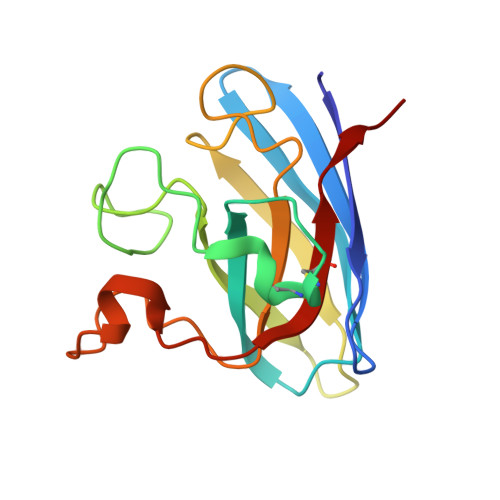
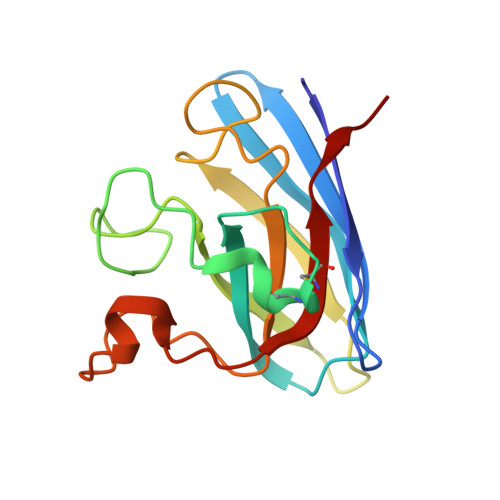
Conformational Variability of the Cu Site in One Subunit of Bovine Cuzn Superoxide Dismutase: The Importance of Mobility in the Glu119-Leu142 Loop Region for Catalytic Function.
Hough, M.A., Strange, R.W., Hasnain, S.S.(2000) J Mol Biology 304: 231
- PubMed: 11080458 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4186
- Primary Citation Related Structures:
1E9O, 1E9P, 1E9Q - PubMed Abstract:
The structure of the catalytic site in one subunit of bovine CuZn superoxide dismutase is shown to be highly variable. A series of crystal structures at approximately 1.7 A have been determined using data collected from different crystals. Several conformations are observed for the copper site from one of the subunits. These conformations lie between those expected for the Cu(II) and Cu(I) forms of the enzyme and may represent a slow positional rearrangement of the Cu site during the crystallisation process due to the presence of a trace reductant in the mother liquor. These states perhaps indicate some functionally relevant structural steps that ultimately result in the breakage of the imidazolate bridge between the two metal sites. This behaviour is not observed for the second subunit of the dimeric enzyme, which remains in the five-coordinate, distorted square planar geometry in all cases. We suggest that this asymmetric behaviour may be caused by the lack of mobility for the Glu119-Leu142 loop region in the second subunit caused by crystal contacts. This region forms one wall of the active-site cavity, and its mobility has been suggested, via molecular dynamics studies, to be important for the catalytic mechanism.
- CLRC Daresbury Laboratory, Daresbury, Warrington, WA4 4AD, UK.
Organizational Affiliation: