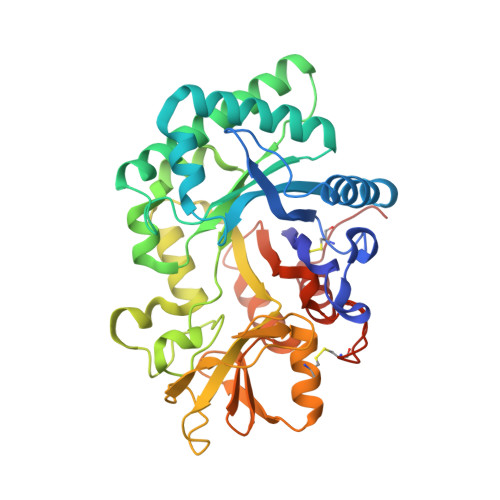
The Crystal Structure of a Novel Mammalian Lectin, Ym1, Suggests a Saccharide Binding Site
Sun, Y.J., Chang, N.C., Hung, S.I., Chang, A.C., Chou, C.C., Hsiao, C.D.(2001) J Biological Chem 276: 17507
- PubMed: 11278670
- DOI: https://doi.org/10.1074/jbc.M010416200
- Primary Citation Related Structures:
1E9L - PubMed Abstract:
Ym1, a secretory protein synthesized by activated murine peritoneal macrophages, is a novel mammalian lectin with a binding specificity to GlcN. Lectins are responsible for carbohydrate recognition and for mediating cell-cell and cell-extracellular matrix interactions in microbes, plants, and animals. Glycosaminoglycan heparin/heparan sulfate binding ability was also detected in Ym1. We report here the three-dimensional structure of Ym1 at 2.5-A resolution by x-ray crystallography. The crystal structure of Ym1 consists of two globular domains, a beta/alpha triose-phosphate isomerase barrel domain and a small alpha + beta folding domain. A notable electron density of sugar is detected in the Ym1 crystal structure. The saccharide is located inside the triose-phosphate isomerase domain at the COOH terminal end of the beta-strands. Both hydrophilic and hydrophobic interactions are noted in the sugar-binding site in Ym1. Despite the fact that Ym1 is not a chitinase, structurally, Ym1 shares significant homology with chitinase A of Serratia marcescens. Ym1 and chitinase A have a similar carbohydrate binding cleft. This study provides new structure information, which will lead to better understanding of the biological significance of Ym1 and its putative gene members.
- Institute of Molecular Biology, Academia Sinica, Taipei, Taiwan 115, Republic of China.
Organizational Affiliation: