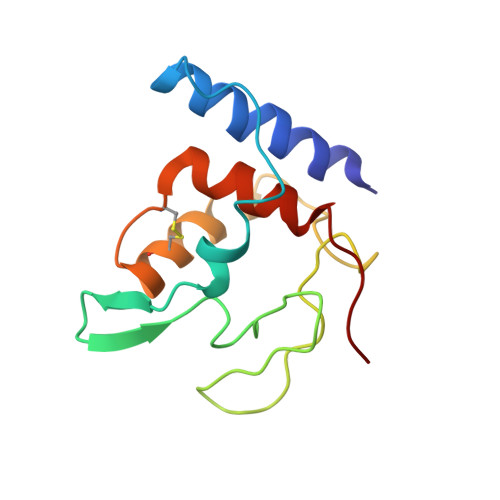
Solution Structure of Methylophilus Methylotrophus Cytochrome C": Insights Into the Structural Basis of Haem-Ligand Detachment
Brennan, L., Turner, D.L., Fareleira, P., Santos, H.(2001) J Mol Biology 308: 353
- PubMed: 11327772 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4600
- Primary Citation Related Structures:
1E8E - PubMed Abstract:
Cytochrome c" from Methylophilus methylotrophus is a monohaem protein with 124 amino acid residues. The iron has two histidine ligands in the oxidised form, one of which detaches and picks up a proton when the protein is reduced. Thus, both forms are paramagnetic. The structure of the oxidised form in solution, determined from NMR data is presented. The family of structures has an average backbone rmsd value of 0.53 A, and a heavy atom rmsd value of 0.95 A, within a target function range of 32 %. This structure is related to class I cytochromes with an additional helix at the N terminus. The haem-binding site occurs in a domain essentially lacking secondary structure motifs and the axial histidinyl residues were found in an unusual near perpendicular orientation. Moreover, a disulfide bridge is present, an uncommon structural feature among c-type cytochromes. The disulfide bridge, linking cysteine residues 96 and 104, forms a loop that confers rigidity and is essential to the detachment of the axial histidine (His95) as demonstrated by chemical disruption of the S-S bond. A route for protonation of the distal histidine involving haem propionate 17 is proposed and discussed in the light of available models for complex membrane proton pumps.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Rua da Quinta Grande, 6 Apt. 127, Oeiras, Portugal.
Organizational Affiliation: