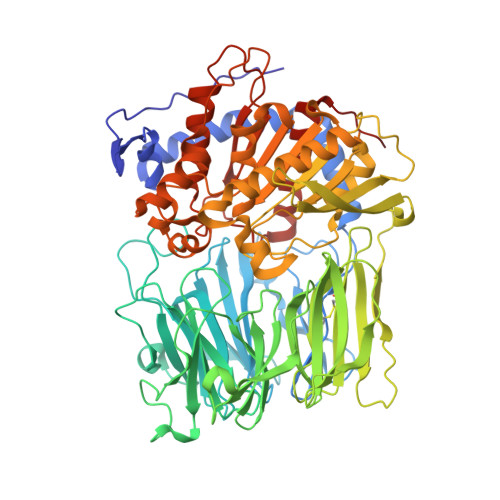
Catalysis of Serine Oligopeptidases is Controlled by a Gating Filter Mechanism
Fulop, V., Szeltner, Z., Polgar, L.(2000) EMBO Rep 1: 277
- PubMed: 11256612 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/embo-reports/kvd048
- Primary Citation Related Structures:
1E5T - PubMed Abstract:
Proteases have a variety of strategies for selecting substrates in order to prevent uncontrolled protein degradation. A recent crystal structure determination of prolyl oligopeptidase has suggested a way for substrate selection involving an unclosed seven-bladed beta-propeller domain. We have engineered a disulfide bond between the first and seventh blades of the propeller, which resulted in the loss of enzymatic activity. These results provided direct evidence for a novel strategy of regulation in which oscillating propeller blades act as a gating filter during catalysis, letting small peptide substrates into the active site while excluding large proteins to prevent accidental proteolysis.
- Department of Biological Sciences, University of Warwick, Coventry, UK.
Organizational Affiliation: