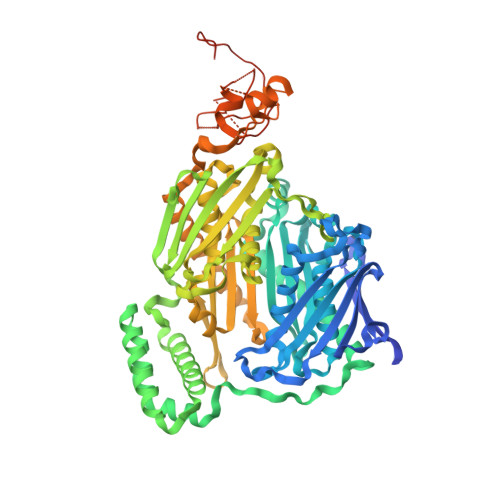
A Duplicated Fold is the Structural Basis for Polynucleotide Phosphorylase Catalytic Activity, Processivity, and Regulation
Symmons, M.F., Jones, G.H., Luisi, B.F.(2000) Structure 8: 1215
- PubMed: 11080643 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00521-9
- Primary Citation Related Structures:
1E3H, 1E3P - PubMed Abstract:
Polynucleotide phosphorylase (PNPase) is a polyribonucleotide nucleotidyl transferase (E.C.2.7.7.8) that degrades mRNA in prokaryotes. Streptomyces antibioticus PNPase also assays as a guanosine 3'-diphosphate 5'-triphosphate (pppGpp) synthetase (E.C.2.7.6.5). It may function to coordinate changes in mRNA lifetimes with pppGpp levels during the Streptomyces lifecycle. The structure of S. antibioticus PNPase without bound RNA but with the phosphate analog tungstate bound at the PNPase catalytic sites was determined by X-ray crystallography and shows a trimeric multidomain protein with a central channel. The structural core has a novel duplicated architecture formed by association of two homologous domains. The tungstate derivative structure reveals the PNPase active site in the second of these core domains. Structure-based sequence analysis suggests that the pppGpp synthetase active site is located in the first core domain. This is the first structure of a PNPase and shows the structural basis for the trimer assembly, the arrangement of accessory RNA binding domains, and the likely catalytic residues of the PNPase active site. A possible function of the trimer channel is as a contribution to both the processivity of degradation and the regulation of PNPase action by RNA structural elements.
- Department of Biochemistry, University of Cambridge, Cambridge, United Kingdom. mfs@mole.bio.cam.ac.uk
Organizational Affiliation: