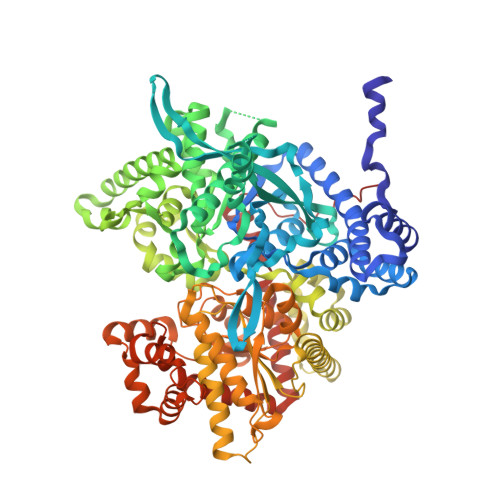
Flavopiridol Inhibits Glycogen Phosphorylase by Binding at the Inhibitor Site
Oikonomakos, N.G., Schnier, J.B., Zographos, S.E., Skamnaki, V.T., Tsitsanou, K.E., Johnson, L.N.(2000) J Biological Chem 275: 34566
- PubMed: 10924512 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M004485200
- Primary Citation Related Structures:
1C8K, 1E1Y, 1GFZ - PubMed Abstract:
Flavopiridol (L86-8275) ((-)-cis-5, 7-dihydroxy-2-(2-chlorophenyl)-8-[4-(3-hydroxy-1-methyl)-piperidinyl] -4H-benzopyran-4-one), a potential antitumor drug, currently in phase II trials, has been shown to be an inhibitor of muscle glycogen phosphorylase (GP) and to cause glycogen accumulation in A549 non-small cell lung carcinoma cells (Kaiser, A., Nishi, K., Gorin, F.A., Walsh, D.A., Bradbury, E. M., and Schnier, J. B., unpublished data). Kinetic experiments reported here show that flavopiridol inhibits GPb with an IC(50) = 15.5 microm. The inhibition is synergistic with glucose resulting in a reduction of IC(50) for flavopiridol to 2.3 microm and mimics the inhibition of caffeine. In order to elucidate the structural basis of inhibition, we determined the structures of GPb complexed with flavopiridol, GPb complexed with caffeine, and GPa complexed with both glucose and flavopiridol at 1.76-, 2.30-, and 2.23-A resolution, and refined to crystallographic R values of 0.216 (R(free) = 0.247), 0.189 (R(free) = 0.219), and 0.195 (R(free) = 0.252), respectively. The structures provide a rational for flavopiridol potency and synergism with glucose inhibitory action. Flavopiridol binds at the allosteric inhibitor site, situated at the entrance to the catalytic site, the site where caffeine binds. Flavopiridol intercalates between the two aromatic rings of Phe(285) and Tyr(613). Both flavopiridol and glucose promote the less active T-state through localization of the closed position of the 280s loop which blocks access to the catalytic site, thereby explaining their synergistic inhibition. The mode of interactions of flavopiridol with GP is different from that of des-chloro-flavopiridol with CDK2, illustrating how different functional parts of the inhibitor can be used to provide specific and potent binding to two different enzymes.
- Institute of Biological Research and Biotechnology, The National Hellenic Research Foundation, 48 Vas. Constantinou Avenue, Athens 11635, Greece. ngo@eie.gr
Organizational Affiliation: