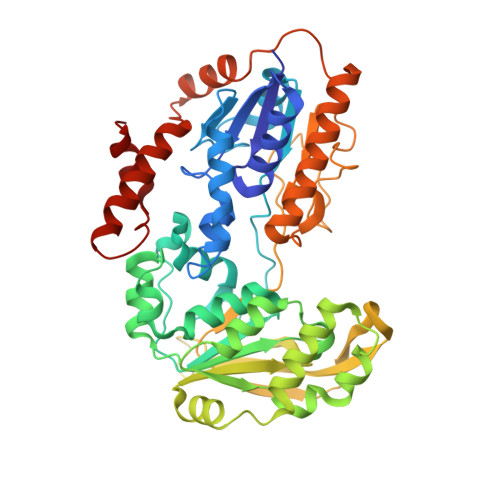
Crystal Structures of Adrenodoxin Reductase in Complex with Nadp+ and Nadph Suggesting a Mechanism for the Electron Transfer of an Enzyme Family
Ziegler, G.A., Schulz, G.E.(2000) Biochemistry 39: 10986
- PubMed: 10998235 Search on PubMed
- DOI: https://doi.org/10.1021/bi000079k
- Primary Citation Related Structures:
1E1K, 1E1L, 1E1M, 1E1N - PubMed Abstract:
Adrenodoxin reductase is a flavoenzyme that shuffles electrons for the biosynthesis of steroids. Its chain topology belongs to the glutathione reductase family of disulfide oxidoreductases, all of which bind FAD at equivalent positions. The three reported structures of adrenodoxin reductase were ligated with reduced and oxidized NADP and have now confirmed this equivalence also for the NADP-binding site. Remarkably, the conformations and relative positions of the prosthetic group FAD and the cofactor NADP have been conserved during protein evolution despite very substantial changes in the polypeptide. The ligated enzymes showed small changes in the domain positions. When compared with the structure of the NADP-free enzyme, these positions correspond to several states of the domain motion during NADP binding. On the basis of the observed structures, we suggest an enzymatic mechanism for the subdivision of the received two-electron package into the two single electrons transferred to the carrier protein adrenodoxin. The data banks contain 10 sequences that are closely related to bovine adrenodoxin reductase. Most of them code for gene products with unknown functions. Within this family, the crucial residues of adrenodoxin reductase are strictly conserved. Moreover, the putative docking site of the carrier is rather well conserved. Five of the family members were assigned names related to ferredoxin:NADP(+) reductase, presumably because adrenodoxin reductase was considered a member of this functionally similar family. Since this is not the case, the data bank entries should be corrected.
- Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, Albertstrasse 21, D-79104 Freiburg im Breisgau, Germany.
Organizational Affiliation: