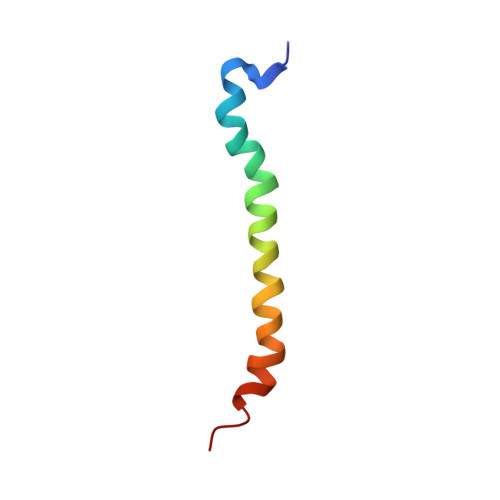
The Solution Structure of Rhodobacter Sphaeroides Lh1 B Reveals Two Helical Domains Separated by a Flexible Region: Structural Consequences for the Lh1 Complex
Conroy, M.J., Westerhuis, W., Parkes-Loach, P.S., Loach, P.A., Hunter, C.N., Williamson, M.P.(2000) J Mol Biology 298: 83
- PubMed: 10756106 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3649
- Primary Citation Related Structures:
1DX7 - PubMed Abstract:
Here, the solution structure of the Rhodobacter sphaeroides core light-harvesting complex beta polypeptide solubilised in chloroform:methanol is presented. The structure, determined by homonuclear NMR spectroscopy and distance geometry, comprises two alpha helical regions (residue -34 to -15 and -11 to +6, using the numbering system in which the conserved histidine residue is numbered zero) joined by a more flexible four amino acid residue linker. The C-terminal helix forms the membrane spanning region in the intact LH1 complex, whilst the N-terminal helix must lie in the lipid head groups or in the cytoplasm, and form the basis of interaction with the alpha polypeptide. The structure of a mutant beta polypeptide W(+9)F was also determined. This mutant, which is deficient in a hydrogen bond donor to the bacteriochlorophyll, showed an identical structure to the wild-type, implying that observed differences in interaction with other LH1 polypeptides must arise from cofactor binding. Using these structures we propose a modification to existing models of the intact LH1 complex by replacing the continuous helix of the beta polypeptide with two helices, one of which lies at an acute angle to the membrane plane. We suggest that a key difference between LH1 and LH2 is that the beta subunit is more bent in LH1. This modification puts the N terminus of LH1beta close to the reaction centre H subunit, and provides a rationale for the different ring sizes of LH1 and LH2 complexes.
- Krebs Institute, Department of Molecular Biology and Biotechnology, University of Sheffield, UK.
Organizational Affiliation: