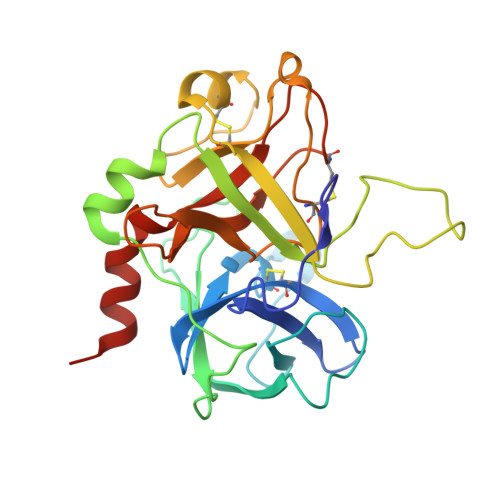
Crystallographic analysis at 3.0-A resolution of the binding to human thrombin of four active site-directed inhibitors.
Banner, D.W., Hadvary, P.(1991) J Biological Chem 266: 20085-20093
- PubMed: 1939071 Search on PubMed
- Primary Citation Related Structures:
1DWB, 1DWC, 1DWD, 1DWE - PubMed Abstract:
The mode of binding of four active-site directed inhibitors to human thrombin has been determined by x-ray crystallographic analysis. The inhibitors studied are benzamidine, PPACK, NAPAP, and MD-805, of which the last three are compounds evolved specifically to inhibit thrombin. Crystal structures were determined in the presence of both the inhibitor and the undecapeptide [des-amino Asp55]hirudin(55-65) which binds distant from the active site. Despite having significantly different chemical structures, NAPAP and MD-805 bind to thrombin in a very similar "inhibitor binding mode" which is not that expected by direct analogy with the binding of substrate. Both inhibitors bind to thrombin in a similar way as to trypsin, but thrombin has an extra loop, the "Tyr-Pro-Pro-Trp loop," not present in trypsin, which gives further binding interactions and is seen to move somewhat to accommodate binding of the different inhibitors. The fact that NAPAP and MD-805 require different stereochemistry for potent inhibition is demonstrated, and its structural basis clarified. The wealth of data on analogs and variants of these lead compounds is shown to be compatible with this inhibitor binding mode.
- Pharmaceutical Research Departments, F. Hoffmann-La Roche Ltd., Basel, Switzerland.
Organizational Affiliation: