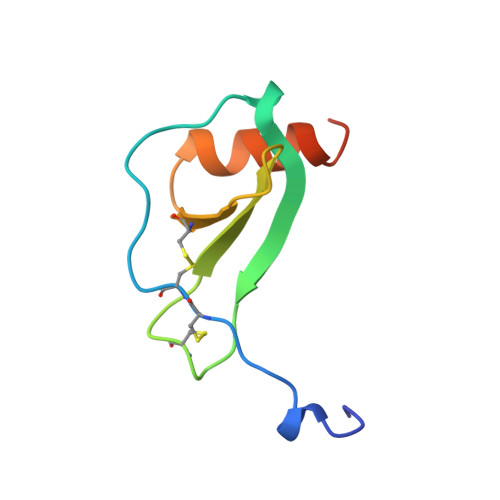
The structure of MCP-1 in two crystal forms provides a rare example of variable quaternary interactions.
Lubkowski, J., Bujacz, G., Boque, L., Domaille, P.J., Handel, T.M., Wlodawer, A.(1997) Nat Struct Biol 4: 64-69
- PubMed: 8989326 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0197-64
- Primary Citation Related Structures:
1DOK, 1DOL - PubMed Abstract:
The X-ray crystal structure of recombinant human monocyte chemoattractant protein (MCP-1) has been solved in two crystal forms. One crystal form (P), refined to 1.85 A resolution, contains a dimer in the asymmetric unit, while the other (I) contains a monomer and was refined at 2.4 A. Although both crystal forms grow together in the same droplet, the respective quaternary structures of the protein differ dramatically. In addition, both X-ray structures differ to a similar extent from the solution structure of MCP-1. Such extent of variability of quaternary structures is unprecedented. In the crystal structures, the well-ordered N termini of MCP-1 form 3(10)-helices. Comparison of the three MCP-1 structures revealed a direct correlation between the main-chain conformation of the first two cysteine residues and the quaternary arrangements. These data can be used to explain the structural basis for the assignment of residues responsible for biological activity.
- Macromolecular Structure Laboratory, NCI-Frederick Cancer Research and Development Center, ABL-Basic Research Program, Maryland 21702, USA.
Organizational Affiliation: