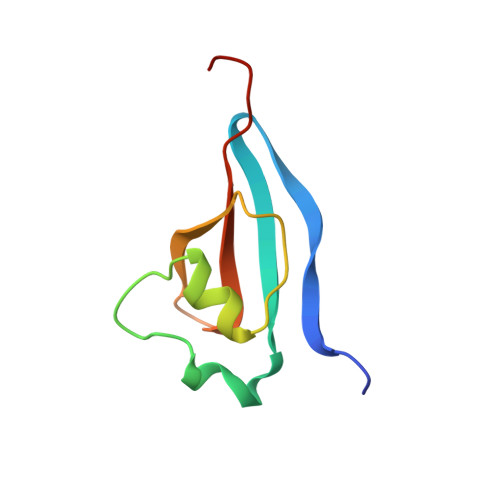
Crystal structure of the human cell cycle protein CksHs1: single domain fold with similarity to kinase N-lobe domain.
Arvai, A.S., Bourne, Y., Hickey, M.J., Tainer, J.A.(1995) J Mol Biology 249: 835-842
- PubMed: 7791211 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1995.0341
- Primary Citation Related Structures:
1DKS, 1DKT - PubMed Abstract:
The structure of the human CksHs1 homolog of the yeast cell-cycle regulatory proteins suc1 and CKS1, which bind to the catalytic subunit of the cyclin-dependent kinases (Cdks) and are essential for yeast cell-cycle progression in vivo, has been determined at 2.9 A resolution. The CksHs1 single polypeptide domain fold, which consists of a four-stranded beta-sheet flanked by two alpha-helices, is dramatically different from the subunit conformation and assembly of the homologous CksHs2, but strikingly similar to the Cdk N-lobe domain fold. The CksHs1 structure identifies sequence-conserved residues Glu61 to His65 as a novel beta-hinge region that folds back to form a beta-hairpin with CksHs1 subunit, whereas this hinge is unfolded to form an extended beta-strand exchange between two CksHs2 subunits. Phosphate and the phosphate analog metavanadate bind CksHs1 in a shallow pocket and interact with five conserved residues (Lys11, Arg20, Ser51, Trp54 and Arg71) suggesting a specific Cks recognition site for a phosphorylated Cdk residue. The dramatic changes to the Cks fold, assembly and exposed conserved surface brought about by switching between the bent and extended hinge conformations are potentially important for the functions of this Cks homolog and could explain conflicting activities inferred from different types of genetic experiments.
- Department of Molecular Biology, Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: