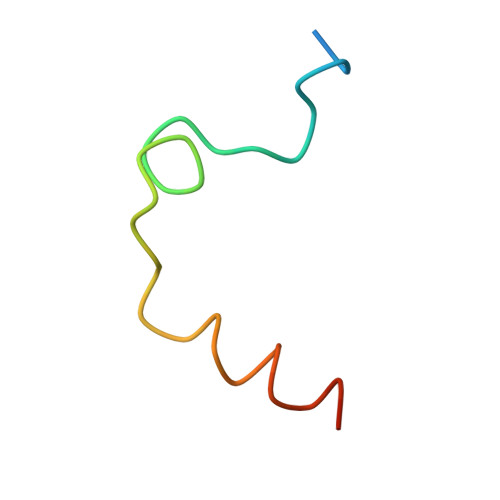
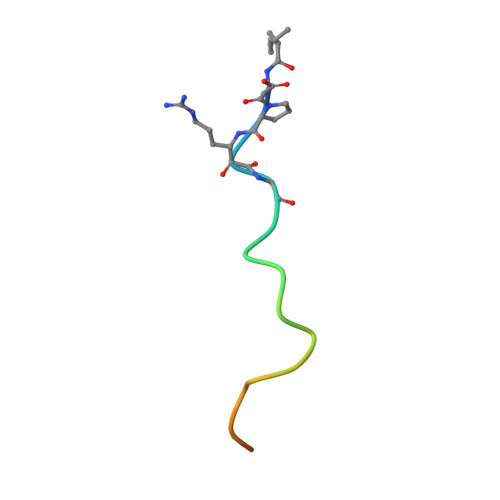
Synthesis, structure, and structure-activity relationships of divalent thrombin inhibitors containing an alpha-keto-amide transition-state mimetic.
Krishnan, R., Tulinsky, A., Vlasuk, G.P., Pearson, D., Vallar, P., Bergum, P., Brunck, T.K., Ripka, W.C.(1996) Protein Sci 5: 422-433
- PubMed: 8868478 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560050303
- Primary Citation Related Structures:
1DIT - PubMed Abstract:
A new class of divalent thrombin inhibitors is described that contains an alpha-keto-amide transition-state mimetic linking an active site binding group and a group that binds to the fibrinogen-binding exosite. The X-ray crystallographic structure of the most potent member of this new class, CVS995, shows many features in common with other divalent thrombin inhibitors and clearly defines the transition-state-like binding of the alpha-keto-amide group. The structure of the active site part of the inhibitor shows a network of water molecules connecting both the side-chain and backbone atoms of thrombin and the inhibitor. Direct peptide analogues of the new transition-state-containing divalent thrombin inhibitors were compared using in vitro assays of thrombin inhibition. There was no direct correlation between the binding constants of the peptides and their alpha-keto-amide counterparts. The most potent alpha-keto-amide inhibitor, CVS995, with a Ki = 1 pM, did not correspond to the most potent divalent peptide and contained a single amino acid deletion in the exosite binding region with respect to the equivalent region of the natural thrombin inhibitor hirudin. The interaction energies of the active site, transition state, and exosite binding regions of these new divalent thrombin inhibitors are not additive.
- Department of Chemistry, Michigan State University, East Lansing 48824-1322, USA.
Organizational Affiliation: