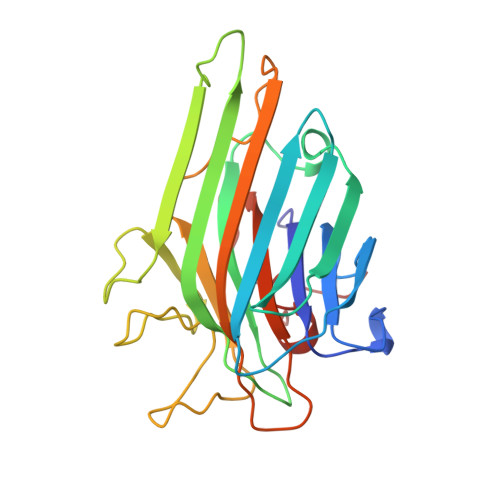
Crystal structure of the lectin from Dioclea grandiflora complexed with core trimannoside of asparagine-linked carbohydrates.
Rozwarski, D.A., Swami, B.M., Brewer, C.F., Sacchettini, J.C.(1998) J Biological Chem 273: 32818-32825
- PubMed: 9830028 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.273.49.32818
- Primary Citation Related Structures:
1DGL - PubMed Abstract:
The seed lectin from Dioclea grandiflora (DGL) has recently been shown to possess high affinity for 3, 6-di-O-(alpha-D-mannopyranosyl)-alpha-D-mannopyranose, the core trimannoside of asparagine-linked carbohydrates, but lower affinity for biantennary complex carbohydrates. In the previous paper, the thermodynamics of DGL binding to deoxy analogs of the core trimannoside and to a biantennary complex carbohydrate were determined by isothermal titration microcalorimetry. The data suggest that DGL recognizes specific hydroxyl groups of the trimannoside similar to that of the jack bean lectin concanavalin A (ConA) (Gupta, D. Dam, T. K., Oscarson, S., and Brewer, C. F. (1997) J. Biol. Chem. 272, 6388-6392). However, the thermodynamics of DGL binding to certain deoxy analogs and to the complex carbohydrate are different from that of ConA. In the present paper, the x-ray crystal structure of DGL complexed to the core trimannoside was determined to a resolution of 2.6 A. The overall structure of the DGL complex is similar to the structure of the ConA-trimannoside complex (Naismith, J. H., and Field, R. A. (1996) J. Biol. Chem. 271, 972-976). The location and conformation of the bound trimannoside as well as its hydrogen-bonding interactions in both complexes are nearly identical. However, differences exist in the location of two loops outside of the respective binding sites containing residues 114-125 and 222-227. The latter residues affect the location of a network of hydrogen-bonded water molecules that interact with the trisaccharide. Differences in the arrangement of ordered water molecules in the binding site and/or protein conformational differences outside of the binding site may account for the differences in the thermodynamics of binding of the two lectins to deoxy analogs of the trimannoside. Molecular modeling studies suggest how DGL discriminates against binding the biantennary complex carbohydrate relative to ConA.
- Department of Biochemistry, Albert Einstein College of Medicine, Bronx, New York 10461, USA.
Organizational Affiliation: