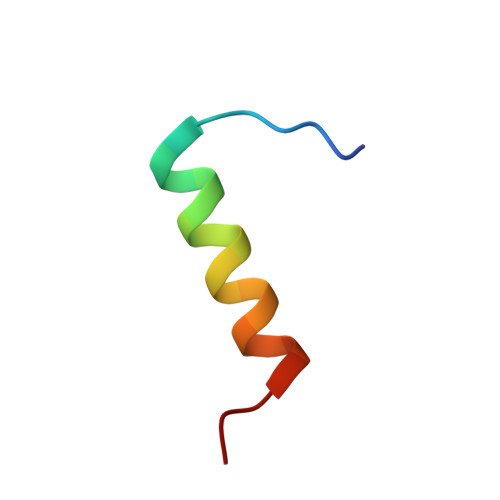
Conformational mapping of the N-terminal segment of surfactant protein B in lipid using 13C-enhanced Fourier transform infrared spectroscopy.
Gordon, L.M., Lee, K.Y., Lipp, M.M., Zasadzinski, J.A., Walther, F.J., Sherman, M.A., Waring, A.J.(2000) J Pept Res 55: 330-347
- PubMed: 10798379 Search on PubMed
- DOI: https://doi.org/10.1034/j.1399-3011.2000.00693.x
- Primary Citation Related Structures:
1DFW - PubMed Abstract:
Synthetic peptides based on the N-terminal domain of human surfactant protein B (SP-B1-25; 25 amino acid residues; NH2-FPIPLPYCWLCRALIKRIQAMIPKG) retain important lung activities of the full-length, 79-residue protein. Here, we used physical techniques to examine the secondary conformation of SP-B1-25 in aqueous, lipid and structure-promoting environments. Circular dichroism and conventional, 12C-Fourier transform infrared (FTIR) spectroscopy each indicated a predominate alpha-helical conformation for SP-B1-25 in phosphate-buffered saline, liposomes of 1-palmitoyl-2-oleoyl phosphatidylglycerol and the structure-promoting solvent hexafluoroisopropanol; FTIR spectra also showed significant beta- and random conformations for peptide in these three environments. In further experiments designed to map secondary structure to specific residues, isotope-enhanced FTIR spectroscopy was performed with 1-palmitoyl-2-oleoyl phosphatidylglycerol liposomes and a suite of SP-B1-25 peptides labeled with 13C-carbonyl groups at either single or multiple sites. Combining these 13C-enhanced FTIR results with energy minimizations and molecular simulations indicated the following model for SP-B1-25 in 1-palmitoyl-2-oleoyl phosphatidylglycerol: beta-sheet (residues 1-6), alpha-helix (residues 8-22) and random (residues 23-25) conformations. Analogous structural motifs are observed in the corresponding homologous N-terminal regions of several proteins that also share the 'saposin-like' (i.e. 5-helix bundle) folding pattern of full-length, human SP-B. In future studies, 13C-enhanced FTIR spectroscopy and energy minimizations may be of general use in defining backbone conformations at amino acid resolution, particularly for peptides or proteins in membrane environments.
- Department of Pediatrics, Martin Luther King, Jr./Drew University Medical Center and UCLA, Los Angeles, CA, USA.
Organizational Affiliation: