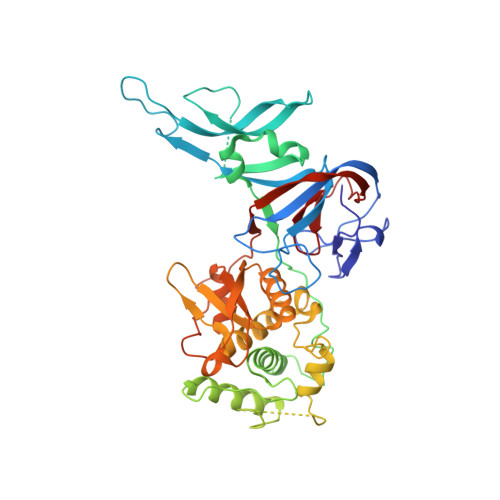
Probing the structure of the PI-SceI-DNA complex by affinity cleavage and affinity photocross-linking.
Hu, D., Crist, M., Duan, X., Quiocho, F.A., Gimble, F.S.(2000) J Biological Chem 275: 2705-2712
- PubMed: 10644733
- DOI: https://doi.org/10.1074/jbc.275.4.2705
- Primary Citation of Related Structures:
1DFA - PubMed Abstract:
The PI-SceI protein is an intein-encoded homing endonuclease that initiates the mobility of its gene by making a double strand break at a single site in the yeast genome. The PI-SceI protein splicing and endonucleolytic active sites are separately located in each of two domains in the PI-SceI structure. To determine the spatial relationship between bases in the PI-SceI recognition sequence and selected PI-SceI amino acids, the PI-SceI-DNA complex was probed by photocross-linking and affinity cleavage methods. Unique solvent-accessible cysteine residues were introduced into the two PI-SceI domains at positions 91, 97, 170, 230, 376, and 378, and the mutant proteins were modified with either 4-azidophenacyl bromide or iron (S)-1-(p-bromoacetamidobenzyl)-ethylenediaminetetraacetate (FeBABE). The phenyl azide-coupled proteins cross-linked to the PI-SceI target sequence, and the FeBABE-modified proteins cleaved the DNA proximal to the derivatized amino acid. The results suggest that an extended beta-hairpin loop in the endonuclease domain that contains residues 376 and 378 contacts the major groove near the PI-SceI cleavage site. Conversely, residues 91, 97, and 170 in the protein splicing domain are in close proximity to a distant region of the substrate. To interpret our results, we used a new PI-SceI structure that is ordered in regions of the protein that bind DNA. The data strongly support a model of the PI-SceI-DNA complex derived from this structure.
- Center for Genome Research, Institute of Biosciences and Technology, Department of Medical Biochemistry, The Texas A & M University System Health Science Center, Houston, Texas 77030, USA.
Organizational Affiliation: