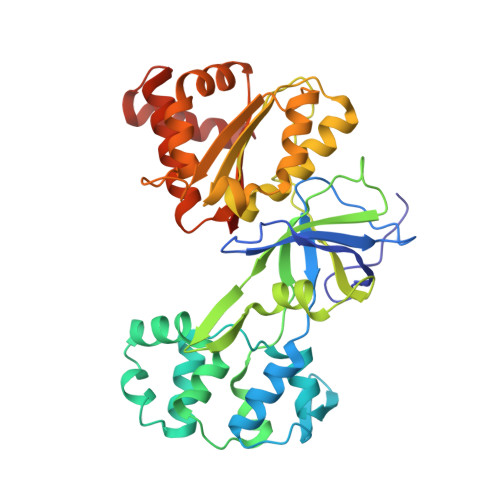
Four crystal structures of the 60 kDa flavoprotein monomer of the sulfite reductase indicate a disordered flavodoxin-like module.
Gruez, A., Pignol, D., Zeghouf, M., Coves, J., Fontecave, M., Ferrer, J.L., Fontecilla-Camps, J.C.(2000) J Mol Biology 299: 199-212
- PubMed: 10860732 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3748
- Primary Citation Related Structures:
1DDG, 1DDI - PubMed Abstract:
Escherichia coli NADPH-sulfite reductase (SiR) is a 780 kDa multimeric hemoflavoprotein composed of eight alpha-subunits (SiR-FP) and four beta-subunits (SiR-HP) that catalyses the six electron reduction of sulfite to sulfide. Each beta-subunit contains a Fe4S4 cluster and a siroheme, and each alpha-subunit binds one FAD and one FMN as prosthetic groups. The FAD gets electrons from NADPH, and the FMN transfers the electrons to the metal centers of the beta-subunit for sulfite reduction. We report here the 1.94 A X-ray structure of SiR-FP60, a recombinant monomeric fragment of SiR-FP that binds both FAD and FMN and retains the catalytic properties of the native protein. The structure can be divided into three domains. The carboxy-terminal part of the enzyme is composed of an antiparallel beta-barrel which binds the FAD, and a variant of the classical pyridine dinucleotide binding fold which binds NADPH. These two domains form the canonic FNR-like module, typical of the ferredoxin NADP+ reductase family. By analogy with the structure of the cytochrome P450 reductase, the third domain, composed of seven alpha-helices, is supposed to connect the FNR-like module to the N-terminal flavodoxine-like module. In four different crystal forms, the FMN-binding module is absent from electron density maps, although mass spectroscopy, amino acid sequencing and activity experiments carried out on dissolved crystals indicate that a functional module is present in the protein. Our results clearly indicate that the interaction between the FNR-like and the FMN-like modules displays lower affinity than in the case of cytochrome P450 reductase. The flexibility of the FMN-binding domain may be related, as observed in the case of cytochrome bc1, to a domain reorganisation in the course of electron transfer. Thus, a movement of the FMN-binding domain relative to the rest of the enzyme may be a requirement for its optimal positioning relative to both the FNR-like module and the beta-subunit.
- Laboratoire de Cristallographie et Cristallogenèse des Protéines Institut de Biologie Structurale J.P. Ebel, CEA-CNRS, Grenoble, France.
Organizational Affiliation: