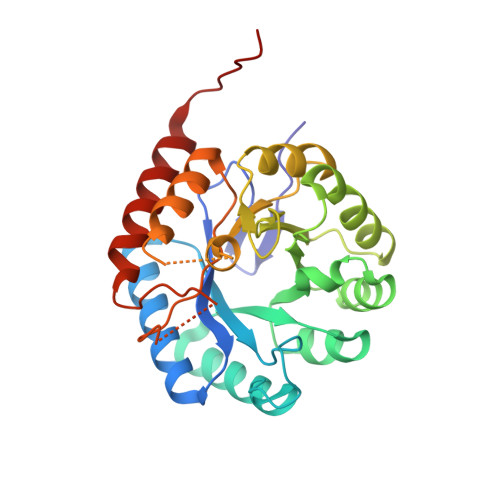
Structure and mechanism of 3-deoxy-D-manno-octulosonate 8-phosphate synthase.
Radaev, S., Dastidar, P., Patel, M., Woodard, R.W., Gatti, D.L.(2000) J Biological Chem 275: 9476-9484
- PubMed: 10734095 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.275.13.9476
- Primary Citation Related Structures:
1D9E - PubMed Abstract:
3-deoxy-D-manno-octulosonate 8-phosphate (KDO8P) synthase catalyzes the condensation of phosphoenolpyruvate (PEP) with arabinose 5-phosphate (A5P) to form KDO8P and inorganic phosphate. KDO8P is the phosphorylated precursor of 3-deoxy-D-manno-octulosonate, an essential sugar of the lipopolysaccharide of Gram-negative bacteria. The crystal structure of the Escherichia coli KDO8P synthase has been determined by multiple wavelength anomalous diffraction and the model has been refined to 2.4 A (R-factor, 19.9%; R-free, 23.9%). KDO8P synthase is a homotetramer in which each monomer has the fold of a (beta/alpha)(8) barrel. On the basis of the features of the active site, PEP and A5P are predicted to bind with their phosphate moieties 13 A apart such that KDO8P synthesis would proceed via a linear intermediate. A reaction similar to KDO8P synthesis, the condensation of phosphoenolpyruvate, and erythrose 4-phosphate to form 3-deoxy-D-arabino-heptulosonate 7-phosphate (DAH7P), is catalyzed by DAH7P synthase. In the active site of DAH7P synthase the two substrates PEP and erythrose 4-phosphate appear to bind in a configuration similar to that proposed for PEP and A5P in the active site of KDO8P synthase. This observation suggests that KDO8P synthase and DAH7P synthase evolved from a common ancestor and that they adopt the same catalytic strategy.
- Department of Biochemistry and Molecular Biology, Wayne State University School of Medicine, Detroit, Michigan 48201, USA.
Organizational Affiliation: