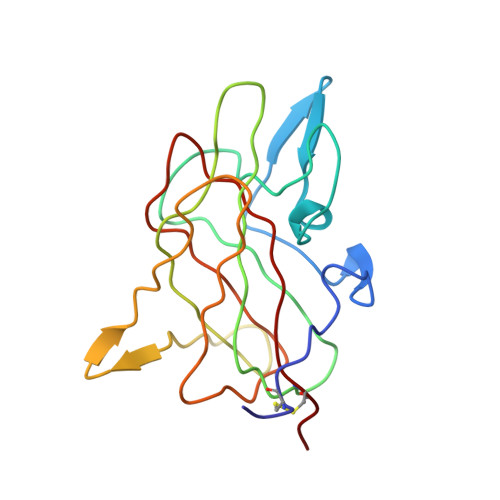
Structure of the C2 domain of human factor VIII at 1.5 A resolution.
Pratt, K.P., Shen, B.W., Takeshima, K., Davie, E.W., Fujikawa, K., Stoddard, B.L.(1999) Nature 402: 439-442
- PubMed: 10586887 Search on PubMed
- DOI: https://doi.org/10.1038/46601
- Primary Citation Related Structures:
1D7P - PubMed Abstract:
Human factor VIII is a plasma glycoprotein that has a critical role in blood coagulation. Factor VIII circulates as a complex with von Willebrand factor. After cleavage by thrombin, factor VIIIa associates with factor IXa at the surface of activated platelets or endothelial cells. This complex activates factor X (refs 6, 7), which in turn converts prothrombin to thrombin in the presence of factor Va (refs 8, 9). The carboxyl-terminal C2 domain of factor VIII contains sites that are essential for its binding to von Willebrand factor and to negatively charged phospholipid surfaces. Here we report the structure of human factor VIII C2 domain at 1.5 A resolution. The structure reveals a beta-sandwich core, from which two beta-turns and a loop display a group of solvent-exposed hydrophobic residues. Behind the hydrophobic surface lies a ring of positively charged residues. This motif suggests a mechanism for membrane binding involving both hydrophobic and electrostatic interactions. The structure explains, in part, mutations in the C2 region of factor VIII that lead to bleeding disorders in haemophilia A.
- Program in Structural Biology, Division of Basic Sciences, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109, USA.
Organizational Affiliation: