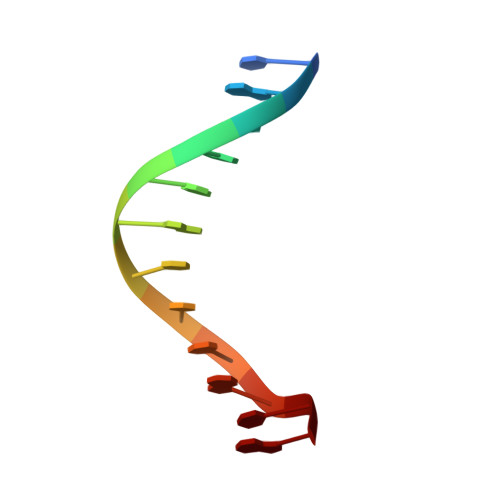
Molecular structure of the B-DNA dodecamer d(CGCAAATTTGCG)2. An examination of propeller twist and minor-groove water structure at 2.2 A resolution.
Edwards, K.J., Brown, D.G., Spink, N., Skelly, J.V., Neidle, S.(1992) J Mol Biology 226: 1161-1173
- PubMed: 1518049 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)91059-x
- Primary Citation Related Structures:
1D65 - PubMed Abstract:
The crystal structure of the dodecanucleotide duplex d(CGCAAATTTGCG)2 has been solved to 2.2 A resolution and refined to an R-factor of 18.1% with the inclusion of 71 water molecules. The structure shows propeller twists of up to -20 degrees for the A.T base-pairs, although there is probably only one (weak) three-centre hydrogen bond in the six base-pair AT narrow minor-groove region. An extensive ribbon of hydration has been located in this groove that has features distinctive from the classic "spine of hydration". Solvation around phosphate groups is described, with several instances of water molecules bridging between phosphates.
- Cancer Research Campaign Biomolecular Structure Unit, Institute of Cancer Research, Sutton, Surrey, U.K.
Organizational Affiliation: