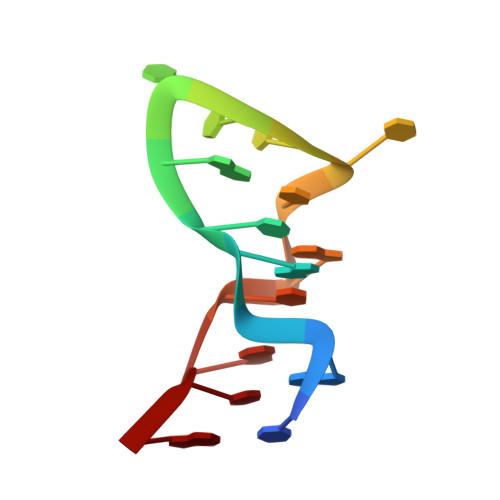
Structure of a T4 hairpin loop on a Z-DNA stem and comparison with A-RNA and B-DNA loops.
Chattopadhyaya, R., Grzeskowiak, K., Dickerson, R.E.(1990) J Mol Biology 211: 189-210
- PubMed: 2299669 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(90)90020-M
- Primary Citation Related Structures:
1D16 - PubMed Abstract:
The synthetic DNA oligomer C-G-C-G-C-G-T-T-T-T-C-G-C-G-C-G crystallizes as a Z-DNA hexamer, capped at one end by a T4 loop. The crystals are monoclinic, space group C2, with a = 57.18 A, b = 21.63 A, c = 36.40 A, beta = 95.22 degrees, and one hairpin molecule per asymmetric unit. The structure of the z-hexamer stem was determined by molecular replacement, and the T4 loop was positioned by difference map methods. The final R factor at 2.1 A resolution for hairpin plus 70 water molecules is 20% for 2 sigma data, with a root-mean-square error of 0.26 A. The (C-G)3 stem resembles the free Z-DNA hexamer with minor crystal packing effects. The T4 loop differs from that observed on a B-DNA stem in solution, or in longer loops in tRNA, in that it shows intraloop and intermolecular interactions rather than base stacking on the final base-pair of the stem. Bases T7, T8 and T9 stack with one another and with the sugar of T7. Two T10 bases from different molecules stack between the C1-G12 terminal base-pairs of a third and fourth molecule, to simulate a T.T "base-pair". Distances between thymine N and O atoms suggest that the two thymine bases are hydrogen bonded, and a keto-enol tautomer pair is favored over disordered keto-keto wobble pairs. The hairpin molecules pack in the crystal in herringbone columns in a manner that accounts well for the observed relative crystal growth rates in a, b and c directions. Hydration seems to be most extensive around the phosphate groups, with lesser hydration within the grooves.
- Department of Chemistry and Biochemistry, Molecular Biology Institute, University of California, Los Angeles 90024.
Organizational Affiliation: