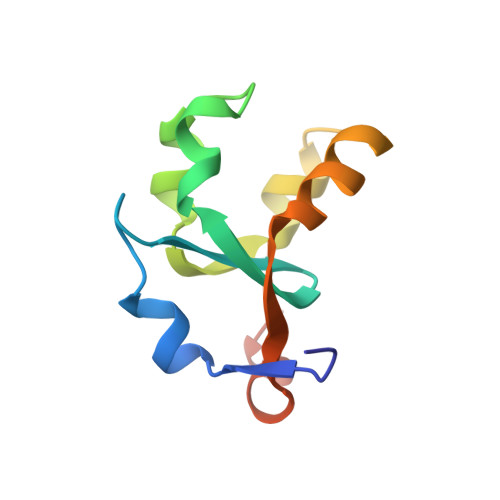
Refinement and structural analysis of bovine cytochrome b5 at 1.5 A resolution.
Durley, R.C., Mathews, F.S.(1996) Acta Crystallogr D Biol Crystallogr 52: 65-76
- PubMed: 15299727 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444995007827
- Primary Citation Related Structures:
1CYO - PubMed Abstract:
The structure of bovine liver cytochrome b(5), a soluble 93-residue proteolytic fragment of a 16 kDa membrane-bound hemoprotein, initially solved at 2.0 A resolution, has been refined at 1.5 A using data collected on a diffractometer. Refinement to 2.0 A resolution used the Hendrickson-Konnert procedure PROLSQ and was then extended to 1.5 A resolution using the program PROFFT. Only residues 3-87 could be identified in the model and these residues together with 93 water molecules gave an agreement factor of R = 0.161 for data in the resolution range 1.5-5 A. The structure was finally refined using the program X-PLOR, which enabled alternate conformers to be modelled for several surface side chains. Residues 1 and 2 at the amino terminus of the protein and residue 88 near the carboxyl terminus could be identified from these electron-density maps. However the remaining disordered carboxy-terminal residues could not successfully be included in the model. A total of 117 solvent molecules were included in the final refinement to give R = 0.164 for the data between 1.5 and 10 A.
- Department of Biochemistry and Molecular Biology, Washington University School of Medicine, St. Louis, MO 63110, USA.
Organizational Affiliation: