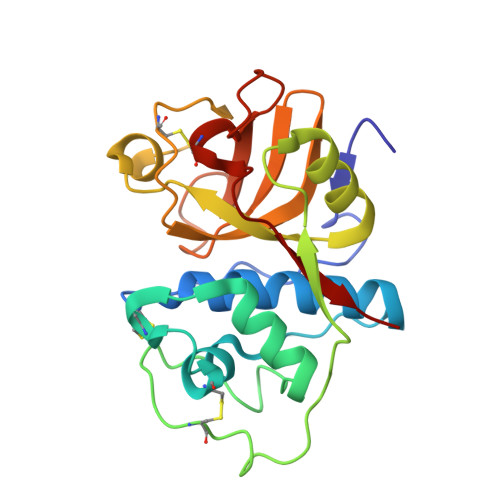
Inhibition mechanism of cathepsin L-specific inhibitors based on the crystal structure of papain-CLIK148 complex.
Tsuge, H., Nishimura, T., Tada, Y., Asao, T., Turk, D., Turk, V., Katunuma, N.(1999) Biochem Biophys Res Commun 266: 411-416
- PubMed: 10600517 Search on PubMed
- DOI: https://doi.org/10.1006/bbrc.1999.1830
- Primary Citation Related Structures:
1CVZ - PubMed Abstract:
Papain was used as an experimental model structure to understand the inhibition mechanism of newly developed specific inhibitors of cathepsin L, the papain superfamily. Recently, we developed a series of cathepsin L-specific inhibitors which are called the CLIK series [(1999) FEBS Lett. 458, 6-10]. Here, we report the complex structure of papain with CLIK148, which is a representative inhibitor from the CLIK series. The inhibitor complex structure was solved at 1.7 A resolution with conventional R 0.177. Unlike other epoxisuccinate inhibitors (E64, CA030, and CA074), CLIK148 uses both prime and nonprime sites, which are important for the specific inhibitory effect on cathepsin L. Also, the specificity for cathepsin L could be explained by the existence of Phe in the P2 site and hydrophobic interaction of N-terminal pyridine ring.
- Institute for Health Sciences, Tokushima Bunri University, Yamashiro-cho, Tokushima, 770-8514, Japan. tsuge@tokushima.bunri-u.ac.jp
Organizational Affiliation: