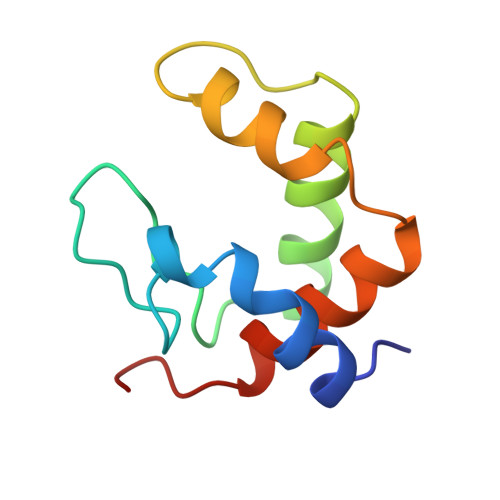
MAD structure of Pseudomonas nautica dimeric cytochrome c552 mimicks the c4 Dihemic cytochrome domain association.
Brown, K., Nurizzo, D., Besson, S., Shepard, W., Moura, J., Moura, I., Tegoni, M., Cambillau, C.(1999) J Mol Biology 289: 1017-1028
- PubMed: 10369779 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2838
- Primary Citation Related Structures:
1CNO - PubMed Abstract:
The monohemic cytochrome c552from Pseudomonas nautica (c552-Pn) is thought to be the electron donor to cytochrome cd1, the so-called nitrite reductase (NiR). It shows as high levels of activity and affinity for the P. nautica NiR (NiR-Pn), as the Pseudomonas aeruginosa enzyme (NiR-Pa). Since cytochrome c552is by far the most abundant electron carrier in the periplasm, it is probably involved in numerous other reactions. Its sequence is related to that of the c type cytochromes, but resembles that of the dihemic c4cytochromes even more closely. The three-dimensional structure of P. nautica cytochrome c552has been solved to 2.2 A resolution using the multiple wavelength anomalous dispersion (MAD) technique, taking advantage of the presence of the eight Fe heme ions in the asymmetric unit. Density modification procedures involving 4-fold non-crystallographic averaging yielded a model with an R -factor value of 17.8 % (Rfree=20.8 %). Cytochrome c552forms a tight dimer in the crystal, and the dimer interface area amounts to 19% of the total cytochrome surface area. Four tighly packed dimers form the eight molecules of the asymmetric unit. The c552dimer is superimposable on each domain of the monomeric cytochrome c4from Pseudomomas stutzeri (c4-Ps), a dihemic cytochrome, and on the dihemic c domain of flavocytochrome c of Chromatium vinosum (Fcd-Cv). The interacting residues which form the dimer are both similar in character and position, which is also true for the propionates. The dimer observed in the crystal also exists in solution. It has been hypothesised that the dihemic c4-Ps may have evolved via monohemic cytochrome c gene duplication followed by evolutionary divergence and the adjunction of a connecting linker. In this process, our dimeric c552structure might be said to constitute a "living fossile" occurring in the course of evolution between the formation of the dimer and the gene duplication and fusion. The availability of the structure of the cytochrome c552-Pn and that of NiR from P. aeruginosa made it possible to identify putative surface patches at which the docking of c552to NiR-Pn may occur.
- Architecture et Fonction des Macromolécules Biologiques, UP R 9039 - CNRS 31, Ch., Joseph Aiguier, Marseille Cedex 20, France.
Organizational Affiliation: