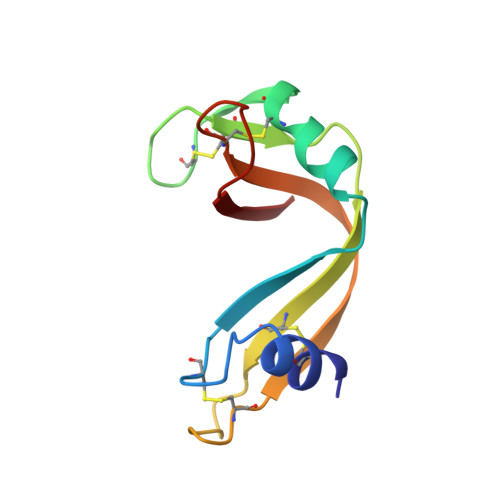
X-ray crystallographic studies of the denaturation of ribonuclease S.
Ratnaparkhi, G.S., Varadarajan, R.(1999) Proteins 36: 282-294
- PubMed: 10409822
- Primary Citation Related Structures:
1CJQ, 1CJR - PubMed Abstract:
In an attempt to view the onset of urea denaturation in ribonuclease we have collected X-ray diffraction data on ribonuclease S crystals soaked in 0, 1.5, 2, 3, and 5 molar urea. At concentrations above 2 M urea, crystals were stabilized by glutaraldehyde crosslinking. We have also collected data on ribonuclease S crystals at low pH in an attempt to study the onset of pH denaturation. The resolution of the datasets range from 1.9 to 3.0 A. Analysis of the structures reveals an increase in disorder with increasing urea concentration. In the 5 M urea structure, this increase in disorder is apparent all over the structure but is larger in loop and helical regions than in the beta strands. The low pH structure shows a very similar pattern of increased disorder. In addition there is a major change in the position of the main chain (> 1 A) in the 65-72 turn region. This region has previously been shown to be involved in one of the initial steps of unfolding in the reduction of ribonuclease A. Crystallographic analyses in the presence of denaturant, when combined with controlled crosslinking, can thus provide detailed structural information that is related to the initial steps of unfolding in solution. Proteins 1999;36:282-294.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India.
Organizational Affiliation: