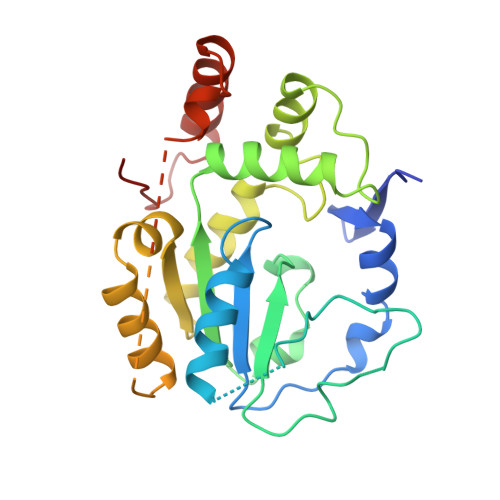
Crystal structure of human catecholamine sulfotransferase.
Bidwell, L.M., McManus, M.E., Gaedigk, A., Kakuta, Y., Negishi, M., Pedersen, L., Martin, J.L.(1999) J Mol Biology 293: 521-530
- PubMed: 10543947 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.3153
- Primary Citation Related Structures:
1CJM - PubMed Abstract:
Sulfonation, like phosphorylation, can modify the activity of a variety of biological molecules. The sulfotransferase enzymes sulfonate neurotransmitters, drugs, steroid hormones, dietary carcinogens and proteins. SULT1A3 specifically sulfonates catecholamines such as dopamine, adrenaline and noradrenaline. The crystal structure of SULT1A3 with a sulfate bound at the active site, has been determined at 2.4 A resolution. Although the core alpha/beta fold is like that of estrogen and heparan sulfotransferases, major differences occur in and around the active site. Most notably, several regions surrounding the active site, including a section of 40 residues, are disordered in SULT1A3. Regions that are topologically equivalent to the disordered parts of SULT1A3 are involved in substrate and cofactor binding in estrogen and heparan sulfotransferase. Flexibility in these regions suggests that ligand binding elicits a disorder-order transition in and around the active site of sulfotransferases and might contribute to the broad substrate specificity of these enzymes.
- Department of Physiology, University of Queensland, Brisbane, Queensland, 4072, Australia.
Organizational Affiliation: