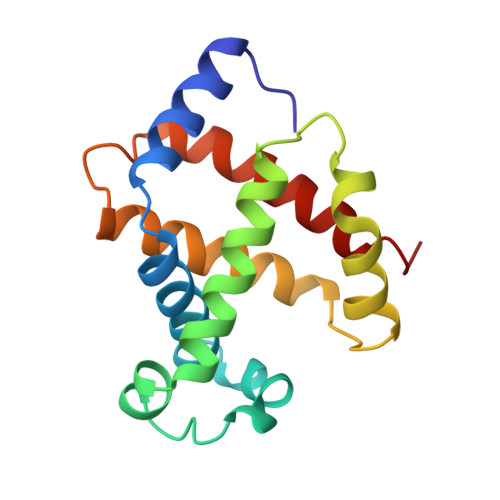
Crystal structure of a protein with an artificial exon-shuffling, module M4-substituted chimera hemoglobin beta alpha, at 2.5 A resolution.
Shirai, T., Fujikake, M., Yamane, T., Inaba, K., Ishimori, K., Morishima, I.(1999) J Mol Biology 287: 369-382
- PubMed: 10080899
- DOI: https://doi.org/10.1006/jmbi.1999.2603
- Primary Citation of Related Structures:
1CH4 - PubMed Abstract:
The crystal structure of the homotetramer of a chimera beta alpha-subunit of human hemoglobin was refined at 2.5 A resolution. The chimera subunit was constructed by replacing an exon-encoded module M4 of the beta-subunit with that of the alpha-subunit, simulating an exon-shuffling event. The implanted module M4 retained the native alpha-subunit structure, while module M3 was disturbed around the site where a new type of intron was recently found. Some of the residues were found in alternative conformations that avoid steric hindrance at the subunit interface. The modules are modestly rigid in their backbone structures by using side-chains to compensate for interface incompatibility.
- Department of Biotechnology and Biomaterial Chemistry Graduate School of Engineering, Nagoya University, Chikusa-Ku, Nagoya, 464-8603, Japan.
Organizational Affiliation: