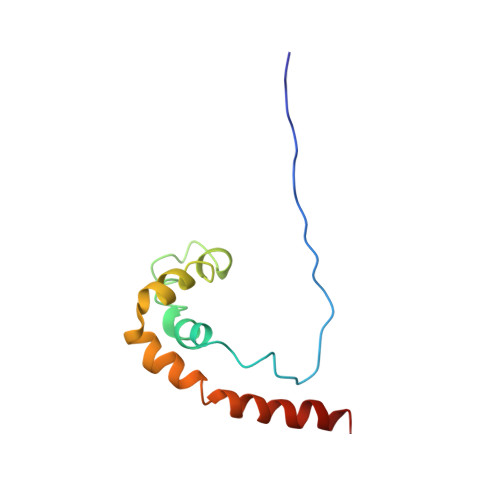
Solution structure of the HMG protein NHP6A and its interaction with DNA reveals the structural determinants for non-sequence-specific binding.
Allain, F.H., Yen, Y.M., Masse, J.E., Schultze, P., Dieckmann, T., Johnson, R.C., Feigon, J.(1999) EMBO J 18: 2563-2579
- PubMed: 10228169 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/18.9.2563
- Primary Citation Related Structures:
1CG7 - PubMed Abstract:
NHP6A is a chromatin-associated protein from Saccharomyces cerevisiae belonging to the HMG1/2 family of non-specific DNA binding proteins. NHP6A has only one HMG DNA binding domain and forms relatively stable complexes with DNA. We have determined the solution structure of NHP6A and constructed an NMR-based model structure of the DNA complex. The free NHP6A folds into an L-shaped three alpha-helix structure, and contains an unstructured 17 amino acid basic tail N-terminal to the HMG box. Intermolecular NOEs assigned between NHP6A and a 15 bp 13C,15N-labeled DNA duplex containing the SRY recognition sequence have positioned the NHP6A HMG domain onto the minor groove of the DNA at a site that is shifted by 1 bp and in reverse orientation from that found in the SRY-DNA complex. In the model structure of the NHP6A-DNA complex, the N-terminal basic tail is wrapped around the major groove in a manner mimicking the C-terminal tail of LEF1. The DNA in the complex is severely distorted and contains two adjacent kinks where side chains of methionine and phenylalanine that are important for bending are inserted. The NHP6A-DNA model structure provides insight into how this class of architectural DNA binding proteins may select preferential binding sites.
- Department of Chemistry and Biochemistry, UCLA, Los Angeles, CA 90095-1569, USA.
Organizational Affiliation: