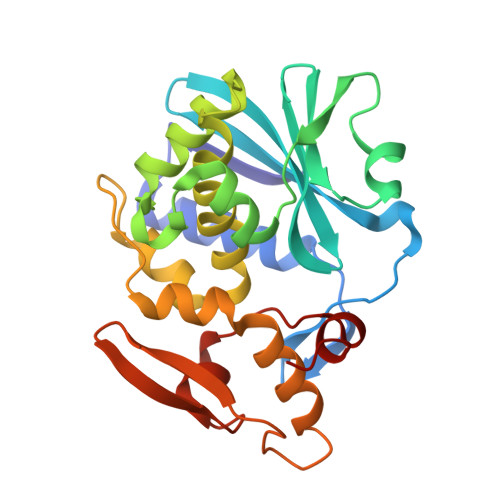
Three-dimensional structure of beta-momorcharin at 2.55 A resolution.
Yuan, Y.R., He, Y.N., Xiong, J.P., Xia, Z.X.(1999) Acta Crystallogr D Biol Crystallogr 55: 1144-1151
- PubMed: 10329776 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444999003297
- Primary Citation Related Structures:
1CF5 - PubMed Abstract:
Beta-Momorcharin (Mr approximately 29 kDa) is a single-chained ribosome-inactivating protein (RIP) with a branched hexasaccharide bound to Asn51. The crystal structure of beta-momorcharin has been determined using the molecular-replacement method and refined to 2. 55 A resolution. The final structural model gave an R factor of 17. 2% and root-mean-square deviations of 0.016 A and 1.76 degrees from ideal bond lengths and bond angles, respectively. beta-Momorcharin contains nine alpha-helices, two 310 helices and three beta-sheets, and its overall structure is similar to those of other single-chained RIPs. Residues Tyr70, Tyr109, Glu158 and Arg161 are expected to define the active site of beta-momorcharin as an rRNA N-glycosidase. The oligosaccharide is linked to the protein through an N-glycosidic bond, beta-GlcNAc-(1-N)-Asn51, and stretches from the surface of the N-terminal domain far from the active site, which suggests that it should not play a role in enzymatic function. The oligosaccharide of each beta-momorcharin molecule interacts with the protein through hydrogen bonds, although in the crystals most of these are intermolecular interactions with the protein atoms in an adjacent unit cell. This is the first example of an RIP structure which provides information about the three-dimensional structure and binding site of the oligosaccharide in the active chains of RIPs.
- Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai 200032, China.
Organizational Affiliation: