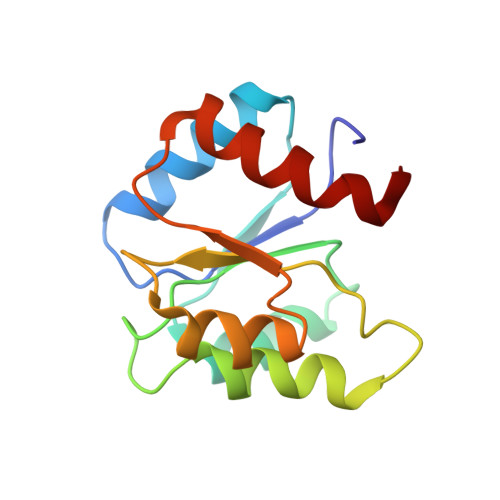
Assignments, secondary structure, global fold, and dynamics of chemotaxis Y protein using three- and four-dimensional heteronuclear (13C,15N) NMR spectroscopy.
Moy, F.J., Lowry, D.F., Matsumura, P., Dahlquist, F.W., Krywko, J.E., Domaille, P.J.(1994) Biochemistry 33: 10731-10742
- PubMed: 8075074 Search on PubMed
- DOI: https://doi.org/10.1021/bi00201a022
- Primary Citation Related Structures:
1CEY - PubMed Abstract:
NMR spectroscopy has been used to study recombinant Escherichia coli CheY, a 128-residue protein involved in regulating bacterial chemotaxis. Heteronuclear three- and four-dimensional (3D and 4D) experiments have provided sequence-specific resonance assignments and quantitation of short-, medium-, and long-range distance restraints from nuclear Overhauser enhancement (NOE) intensities. These distance restraints were further supplemented with measurements of three-bond scalar coupling constants to define the local dihedral angles, and with the identification of amide protons undergoing slow solvent exchange from which hydrogen-bonding patterns were identified. The current model structure shows the same global fold of CheY as existing X-ray structures (Volz & Matsumura, 1991; Stock et al. 1993) with a (beta/alpha)5 motif of five parallel beta-strands at the central core surrounded by three alpha-helices on one face and with two on the opposite side. Heteronuclear 15N-1H relaxation experiments are interpreted to show portions of the protein structure in the Mg2+ binding loop are ill-defined because of slow motion (chemical exchange) on the NMR time scale. Moreover, the presence of Mg2+ disrupts the salt bridge between the highly conserved Lys-109 and Asp-57, the site of phosphorylation.
- DuPont Merck Pharmaceutical Company, Wilmington, Delaware 19880-0328.
Organizational Affiliation: