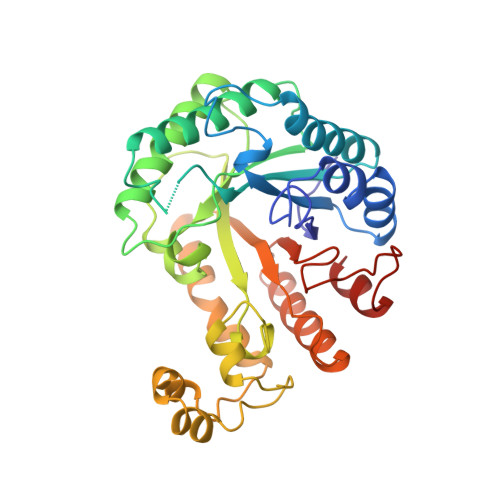
The crystal structure of a family 5 endoglucanase mutant in complexed and uncomplexed forms reveals an induced fit activation mechanism.
Dominguez, R., Souchon, H., Lascombe, M., Alzari, P.M.(1996) J Mol Biology 257: 1042-1051
- PubMed: 8632467 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0222
- Primary Citation Related Structures:
1CEN, 1CEO - PubMed Abstract:
The structures of the Glu140-->Gln mutant of the Clostridium thermocellum endoglucanase CelC in unliganded form (CelC(E140Q)) and in complex with cellohexaose (CelC(E140Q)-Gl(C6)) have been refined to crystallographic R-factors of 19.4% at 1.9 A and 17.8% at 2.3 A resolution, respectively. The structure of CelC(E140Q)-Gl(C6) complex shows two D-glucosyl residues bound to the non-reducing end of the substrate-binding cleft. Comparison of the unliganded and complexes structures reveals conformational changes due to substrate binding, including a significant reorientation of the loop 138-141 which carries the general acid/base catalyst Glu140 in wild-type CelC. Endoglucanase CelC, a family 5 glycohydrolase, exhibits a (beta/alpha)8-fold with an additional subdomain of 54 amino acids inserted between beta-strand 6 and alpha-helix 6. Seven amino acid residues (Arg46, His90, Asn139, Glu140, His198, Tyr200, and Glu280) located close to the catalytic reaction center are strictly conserved in family 5 cellulases. Only three of these residues (His90, Gln140 and Glu280) make direct contacts with the substrate, but all participate in a network of hydrogen bonds which contribute to the stability of the active site architecture and may influence the protonation state of the two catalytic residues. Residue Trp313, which interacts with the nucleophile Glu280 and is within hydrogen bonding distance of the substrate, is involved in a non-proline cis-peptide bond. An aromatic residue occurs at an equivalent position in many other (beta/alpha)8-barrel glycosidases; the presence of a cis-peptide bond at this position in the structures of family 1 beta-glucosidases, family 2 beta-galactosidases, family 5 cellulases, family 17 beta-glucanases, and family 18 chitinases provides further evidence of an evolutionary relationship between glycosyl hydrolases with a (beta/alpha)8- architecture.
- Unité d'Immunologie Structurale, Institut Pasteur, Paris, France.
Organizational Affiliation: