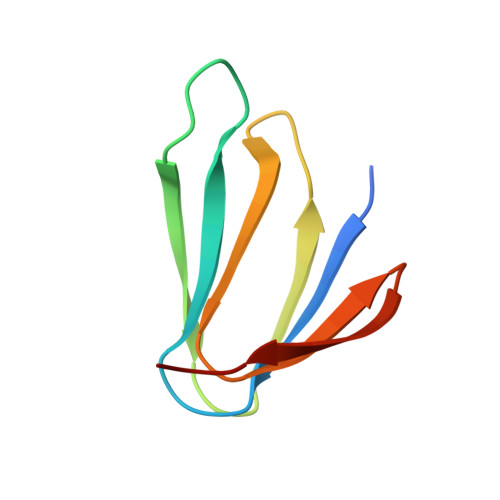
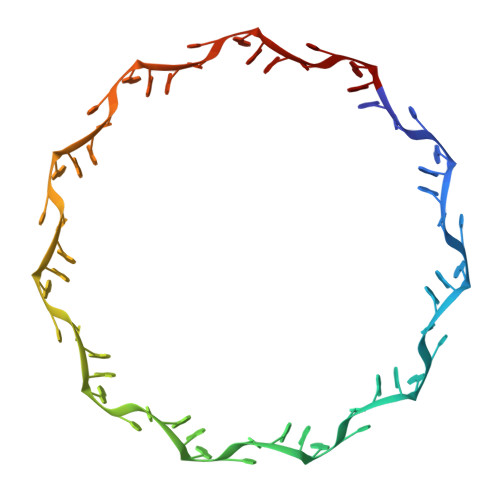
Structure of the trp RNA-binding attenuation protein, TRAP, bound to RNA.
Antson, A.A., Dodson, E.J., Dodson, G., Greaves, R.B., Chen, X., Gollnick, P.(1999) Nature 401: 235-242
- PubMed: 10499579 Search on PubMed
- DOI: https://doi.org/10.1038/45730
- Primary Citation Related Structures:
1C9S - PubMed Abstract:
The trp RNA-binding attenuation protein (TRAP) regulates expression of the tryptophan biosynthetic genes of several bacilli by binding single-stranded RNA. The binding sequence is composed of eleven triplet repeats, predominantly GAG, separated by two or three non-conserved nucleotides. Here we present the crystal structure of a complex of TRAP and a 53-base single-stranded RNA containing eleven GAG triplets, revealing that each triplet is accommodated in a binding pocket formed by beta-strands. In the complex, the RNA has an extended structure without any base-pairing and binds to the protein mostly by specific protein-base interactions. Eleven binding pockets on the circular TRAP 11-mer form a belt with a diameter of about 80 A. This simple but elegant mechanism of arresting the RNA segment by encircling it around a protein disk is applicable to both transcription, when TRAP binds the nascent RNA, and to translation, when TRAP binds the same sequence within a non-coding leader region of the messenger RNA.
- York Structural Biology Laboratory, Department of Chemistry, University of York, UK. fred@yorvic.york.ac.uk
Organizational Affiliation: