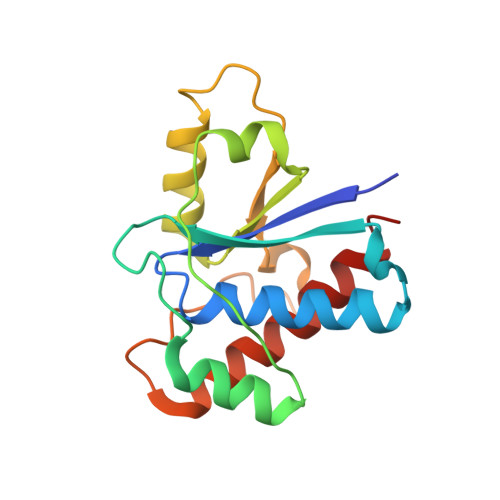
The structure of the bovine protein tyrosine phosphatase dimer reveals a potential self-regulation mechanism.
Tabernero, L., Evans, B.N., Tishmack, P.A., Van Etten, R.L., Stauffacher, C.V.(1999) Biochemistry 38: 11651-11658
- PubMed: 10512620 Search on PubMed
- DOI: https://doi.org/10.1021/bi990381x
- Primary Citation Related Structures:
1C0E - PubMed Abstract:
The bovine protein tyrosine phosphatase (BPTP) is a member of the class of low-molecular weight protein tyrosine phosphatases (PTPases) found to be ubiquitous in mammalian cells. The catalytic site of BPTP contains a CX(5)R(S/T) phosphate-binding motif or P-loop (residues 12-19) which is the signature sequence for all PTPases. Ser19, the final residue of the P-loop motif, interacts with the catalytic Cys12 and participates in stabilizing the conformation of the active site through interactions with Asn15, also in the P-loop. Mutations at Ser19 result in an enzyme with altered kinetic properties with changes in the pK(a) of the neighboring His72. The X-ray structure of the S19A mutant enzyme shows that the general conformation of the P-loop is preserved. However, changes in the loop containing His72 result in a displacement of the His72 side chain that may explain the shift in the pK(a). In addition, it was found that in the crystal, the protein forms a dimer in which Tyr131 and Tyr132 from one monomer insert into the active site of the other monomer, suggesting a dual-tyrosine motif on target sites for this enzyme. Since the activity of this PTPase is reportedly regulated by phosphorylation at Tyr131 and Tyr132, the structure of this dimer may provide a model of a self-regulation mechanism for the low-molecular weight PTPases.
- Department of Biological Sciences, Purdue University, West Lafayette, Indiana 47907, USA.
Organizational Affiliation: