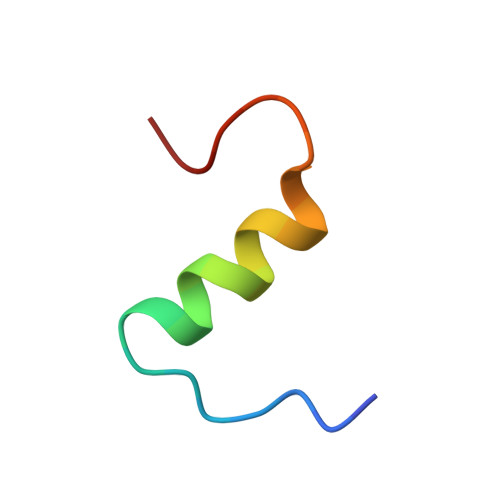
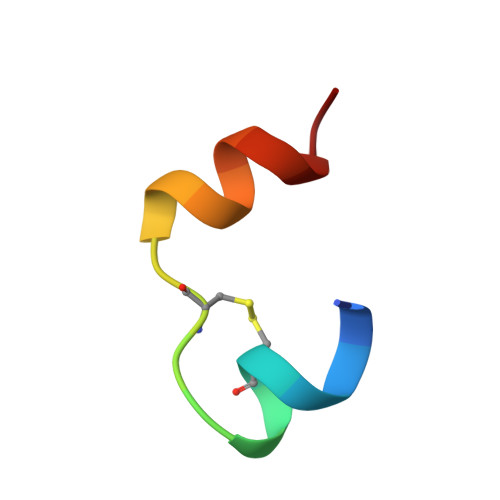The solution structure of a superpotent B-chain-shortened single-replacement insulin analogue.
Kurapkat, G., Siedentop, M., Gattner, H.G., Hagelstein, M., Brandenburg, D., Grotzinger, J., Wollmer, A.(1999) Protein Sci 8: 499-508
- PubMed: 10091652 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.8.3.499
- Primary Citation Related Structures:
1BZV - PubMed Abstract:
This paper reports on an insulin analogue with 12.5-fold receptor affinity, the highest increase observed for a single replacement, and on its solution structure, determined by NMR spectroscopy. The analogue is [D-AlaB26]des-(B27-B30)-tetrapeptide-insulin-B26-amide. C-terminal truncation of the B-chain by four (or five) residues is known not to affect the functional properties of insulin, provided the new carboxylate charge is neutralized. As opposed to the dramatic increase in receptor affinity caused by the substitution of D-Ala for the wild-type residue TyrB26 in the truncated molecule, this very substitution reduces it to only 18% of that of the wild-type hormone when the B-chain is present in full length. The insulin molecule in solution is visualized as an ensemble of conformers interrelated by a dynamic equilibrium. The question is whether the "active" conformation of the hormone, sought after in innumerable structure/function studies, is or is not included in the accessible conformational space, so that it could be adopted also in the absence of the receptor. If there were any chance for the active conformation, or at least a predisposed state to be populated to a detectable extent, this chance should be best in the case of a superpotent analogue. This was the motivation for the determination of the three-dimensional structure of [D-AlaB26]des-(B27-B30)-tetrapeptide-insulin-B26-amide. However, neither the NMR data nor CD spectroscopic comparison of a number of related analogues provided a clue concerning structural features predisposing insulin to high receptor affinity. After the present study it seems more likely than before that insulin will adopt its active conformation only when exposed to the force field of the receptor surface.
- Institut für Biochemie, Rheinisch-Westfälische Technische Hochschule Aachen, Germany.
Organizational Affiliation: